An Open Source, Collaborative Electronic Notebook for Undergraduate Laboratory Classes
Published online:
Abstract
To overcome many of the limitations of the paper laboratory notebook (PLN), we developed an electronic laboratory notebook (e-LN) for the Dynamic Genome course at the University of California. The Dynamic Genome course is an authentic research experience that generates mostly digital data, which is difficult to collect or analyze in a PLN. The e-LN uses free, open-source software that can be accessed through web browsers on either computers or tablet devices. Digital data, such as gel images, spectrophotometer data, screen captures, and bioinformatics data, can be easily recorded in the e-LN. Online access had unexpected pedagogical benefits to both instructors and students. For example, e-LN entries and assignments can be graded in a timely and collaborative manner. Students reported that the templates and feedback were easier with the e-LN than the PLN. Instructors reported that the format improved the quality of scientific record keeping and maximized the usefulness of class time.
Citation
Robb, S., Burnette III, J.M., Chapovskya, A., Palmer, K. and Wessler, S.R. 2015. An Open Source, Collaborative Electronic Notebook for Undergraduate Laboratory Classes. CourseSource. https://doi.org/10.24918/cs.2015.1Article Context
Course
Article Type
Course Level
Vision and Change Core Competencies
Class Type
Audience
Lesson Length
Pedagogical Approaches
Principles of How People Learn
Assessment Type
INTRODUCTION
The Dynamic Genome (DG) course is an authentic research experience in molecular biological and genomic research for first-year undergraduates (1). During the first half of the course students learn key biological concepts such as genetic information transfer and genome variation, while mastering routine laboratory skills including DNA extraction, PCR and gel electrophoresis. Bioinformatics is a major emphasis of the course and students learn to access DNA sequence databases to develop hypotheses and design experiments. The second half of the course consists of a group project in which students generate original data. Because much of these data are in electronic format (e.g. DNA sequence alignments, computational biology software, spectrometric measurements, agarose gel images), the traditional paper laboratory notebook (PLN) was inadequate for recording and analyzing data. Consequently, we sought an alternative laboratory notebook format.
An authentic research course requires accurate and timely record keeping. In the DG course, students prepare a notebook entry before class, record data and take notes during class, and complete data analysis after class. During the first five years of the course, records were kept in a PLN. However, several shortcomings were noted. First, the PLN creates an artificial divide between computational data and laboratory generated data. Second, use of the PLN prolonged the grading cycle and students did not receive timely feedback on pre-lab write-ups and post-lab data analysis. As a result, students would be unprepared for class and repeated the same mistakes. Third and most important, the PLN did not support collaboration between the instructor and student, or between students.
To solve the shortcomings of the PLN we pursued a no-cost solution that was completely paperless, open-source, platform independent, and supported collaboration among students and instructors. A search of existing e-LNs revealed that the vast majority were developed for use in research laboratories and none of these supported broad collaborations (Table 1). Available commercial solutions such as Evernote (2), Blackboard (4), Findings (Findings Software, www.findsapp.com), Accelrys Paperless Lab (Biovia, San Diego, CA) and PerkinElmer E-Notebook (Perkin Elmer, Waltham, MA) require a fee either from the university or the student and do not support student-to-student collaboration. Finally the e-LN must comply with the regulations of the Federal Educational Records Protection Act (FERPA) to protect student rights and privacy (20 U.S.C. ? 1232g; 34 CFR Part 99, http://www2.ed.gov/policy/gen/guid/fpco/ferpa/index.html). Of the existing solutions, only Blackboard and Sakai comply with FERPA regulations. Because we could not find an e-LN that met all of our requirements, we developed the e-LN described in this report.
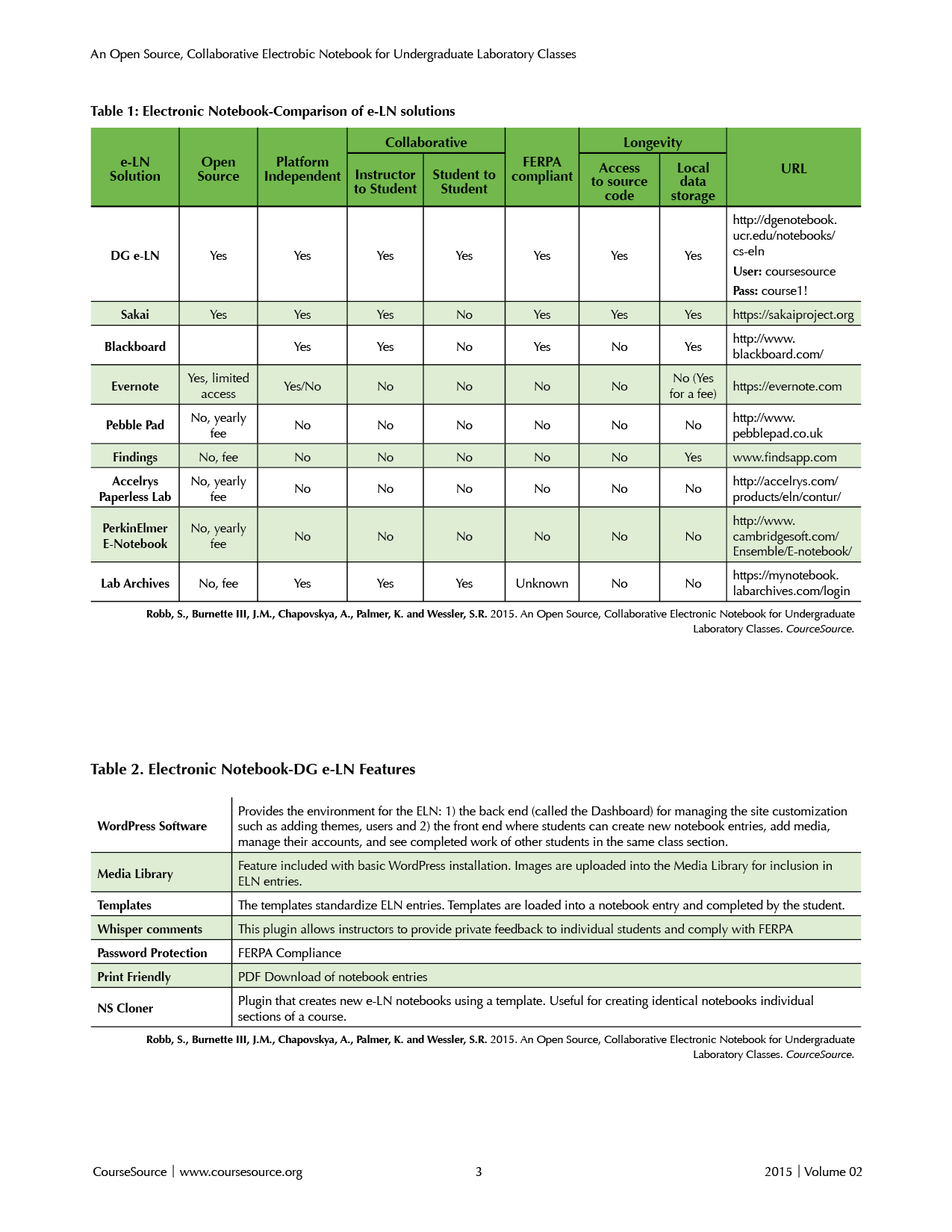
Table 1. Comparison of e-LN Solutions.
Table 2. DG e-LN Features.
IMPLEMENTATION
The e-LN is an implementation of the highly collaborative open-source blogging software WordPress (www.wordpress.org), with data stored using the open-source MySQL database management system (www.mysql.com). Both WordPress and MySQL are free software, are compatible with common operating systems (Mac OS X, Windows, and Linux), and can be installed on university owned computers. Students access the e-LN from any browser-enabled device with Internet access. Installation of both WordPress and MySQL on university computers allows instructors to control access and safeguard educational material as required by FERPA. Finally, both WordPress and MySQL can easily scale from hundreds to thousands of users without additional work or fees. The DG e-LN was developed and deployed for the classroom on an Apple Mac Mini computer running OS X Server and Time Machine backup software.
Table 2 summarizes the key features of the DG e-LN and detailed instructions for installation and configuration can be found at the DG e-LN website (http://dgenotebook.ucr.edu/notebooks/eln-install). WordPress is highly customizable with the use of hundreds of free WordPress applications called "plugins". We installed the basic software and enabled the "Multisite" option to allow individual e-LNs to be set up for every section of a course. Use of this option allows WordPress to manage many notebooks at once so that students and instructors in one lab section can collaborate. The Wordpress "media library" stores uploaded images so that students can easily locate their data and compare it with the data of other students.
All e-LNs have a front page (Figure 1A and see example course link in Table 1) that contains course material required by all students. Instructors access the plugins through the Dashboard (Figure 1B). A very useful plugin that simplifies the enrollment process is the "Add Multiple Users" or "AMU" which allows an instructor to add e-LN users, by uploading a spreadsheet of names and passwords. In this way, a group of students and their teaching assistant can easily collaborate via their e-LN entries. The "Easy Content Templates" plugin provides instructors with the ability to create templates for each notebook entry, by simply clicking on the "Add New" option of the Template link (Figure 1B, blue arrow). The word processor-like editing window makes it easy to enter and format the template as needed. Templates guide students to include the necessary information into a notebook entry and also streamline the grading process. The "Whisper Comments for Class" allows private communication between instructors (or TAs) and students. Thus, a comment about the quality of a student entry or about grades can be discretely discussed and remains private, even after the student's entry is public. Finally, students can download a PDF version of a notebook entry by using the Print Friendly button that appears at the end of a published post.
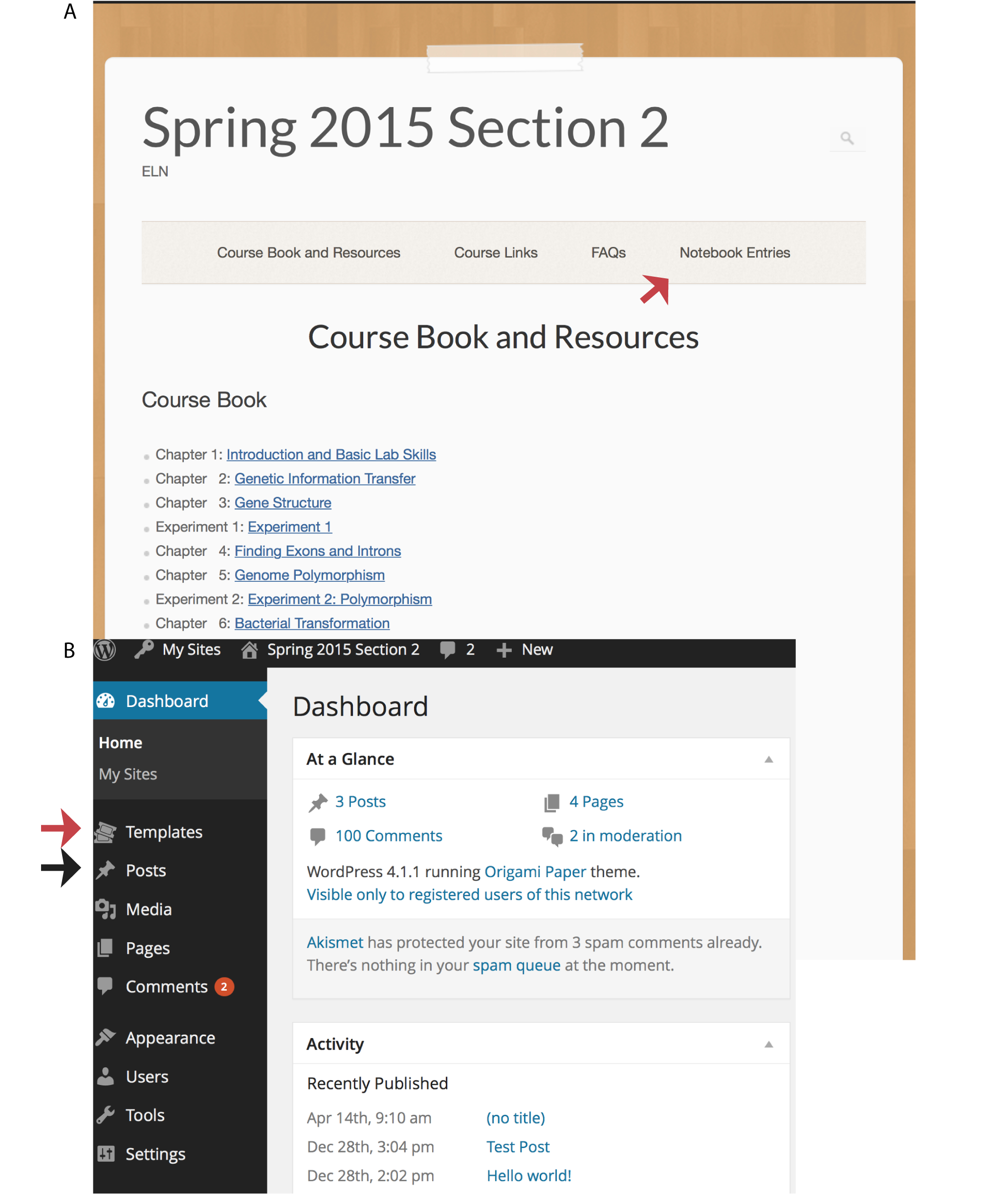
Figure 1. Front page of the e-LN and instructor’s dashboard. A. Notebook entries are accessed using the “Notebook Entries” (red arrow) link at the top. The Course Book chapters and experiments are available in PDF format by clicking on the links. B. The instructor’s dashboard. Links in the black area on the left allow administrative access to areas such as posts (black arrow) and templates (red arrow). See text for details.
Example e-LN Entry
To illustrate how students use the e-LN, we describe an actual class experiment. In this experiment (1), students discover DNA polymorphism among different strains of maize. The complete experiment requires two 3-hour class periods. On Day 1, students discuss experimental design and the PCR technique, learn how to set up PCR samples, and pour agarose gels. On Day 2, students separate and visualize the PCR fragments on gels, photograph their results, and analyze the data. They also learn how to use Multiple Sequence Alignment (MSA) software such as MUSCLE (EBI: http://www.ebi.ac.uk/Tools/msa/muscle/) to identify polymorphisms.
Prior to Day 1, students read the course material, including the protocol for PCR set up and gel preparation, and start their notebook entry by writing the protocol in their own words (see Table 3).

Table 3. Flow of an e-LN post.
Students using a PLN refer to notebook entry instructions about the proper format and content of the pre-lab entry. In contrast, students using the e-LN start a new post (Figure 2A), load the experiment template (Figure 2B and C), enter the requested information (Figure 2D), and submit the completed template to the instructors for comments and grading prior to the start of class. Because all of the preparation and pre-grading occurs prior to class with the e-LN, students are able to correct mistakes and are ready to perform the experiment when they enter the lab. In contrast, instructors first saw student entries in the PLNs during class, requiring significant time to grade, as well as to identify and discuss common misunderstandings. Clearly, the e-LN approach allows students to hit the ground running when they enter the lab, freeing up valuable instructor/TA time to address more important questions about experimental design and protocols.
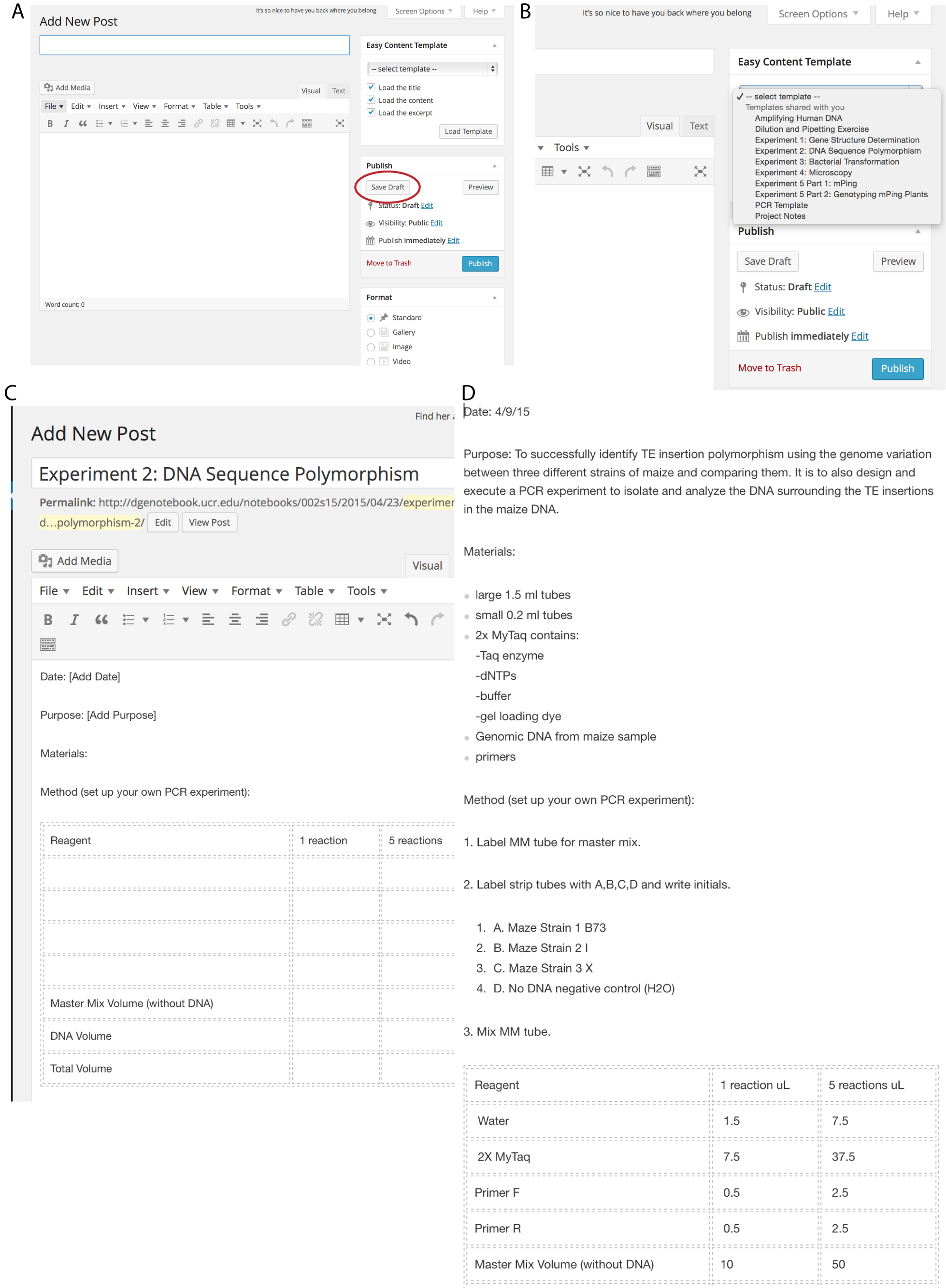
Figure 2. Students are guided through a new notebook entry. A. Start entry by clicking on a new Post in student dashboard. B. Select the template. C. Loaded template (text only shown) is ready to fill out. D. Completed template (text only) is ready to submit to instructor prior to class by clicking the “Save Draft” button (red circle in A).
In the DG classroom/laboratory, students access their e-LN entries using a tablet computer (Figure 3A). We provide Apple iPads for student use in class, although any tablet or other computer can support the e-LN. While students are in class, the notebook entries are in draft phase so students can edit their entries as needed and include notes about their samples or experimental conditions.
On Day 2 of the Polymorphism experiment, students use the camera in the tablet computers to capture gel images via the Fotodyne Discovery/Doc System (Figure 3B, Fotodyne Hartland, WI). Prior to photographing their gels, students place a piece of masking tape on the bottom of the gel, with their names with a black permanent marker (e.g., Sharpie, VWR Lab Markers). The marker ink is visible under UV light, enabling students to locate their images in the e-LN Media Library (included in WordPress). Students can annotate their images or during class or at home using free image editing software, such as YouDoodle (youdoodle.net for the iPad) or Google Drawings (docs.google.com/drawings) (Figure 3C). Then, they can easily upload the annotated image into their e-LN.
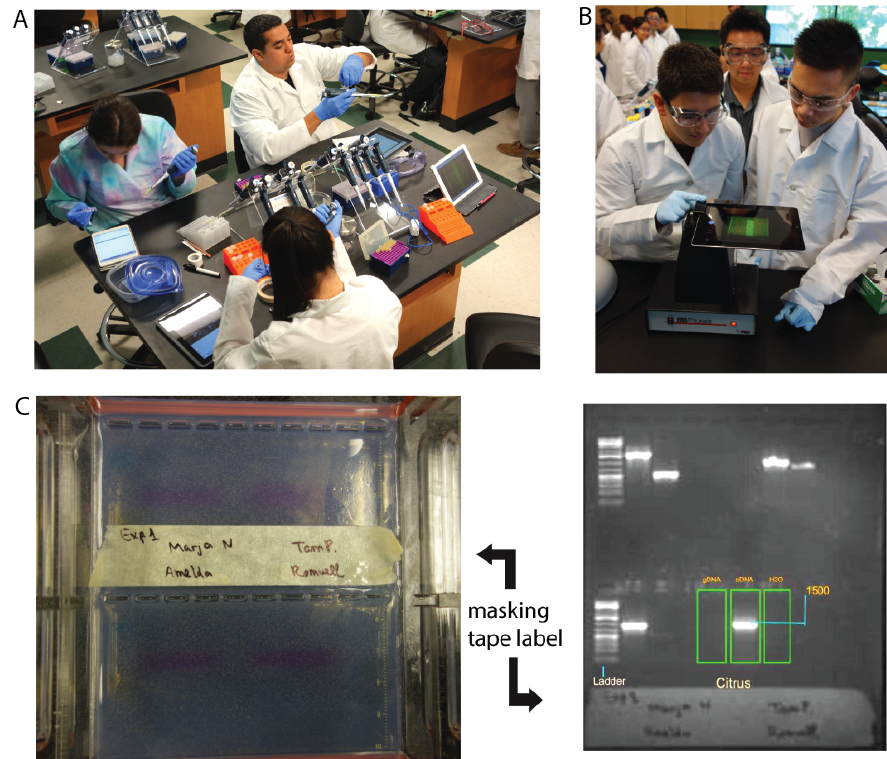

Figure 3. Using the e-LN in class. A. Students access the e-LN at the lab bench. B. Gel image capture using a tablet computer and FotoDyne Fotodyne Discovery/Doc System. C. Annotated image uploaded to post (red arrow).
In contrast, documentation of gel images in a PLN usually involves expensive gel documentation systems that print images onto thermal paper. The printed images were sometimes lost when students forgot to tape them into their notebooks. We find that the use of the WordPress Media Library and the e-LN has greatly improved the quality of image annotation and decreased the loss of gel data.
Computational analysis is an integral part of the DG course. We require students to use bioinformatics software and generate electronic data including GenBank files, DNA alignments, PCR primer design output, and large spreadsheets. Incorporating files of these types is relatively straight forward in the e-LN, but not in the PLN. During Day 2 of the Polymorphism experiment, students use online MSA software (EBI: http://www.ebi.ac.uk/Tools/msa/muscle/) to compare the DNA sequences of two PCR products that differ by an insertion of a transposable element. The sequences, provided by the instructors, can be accessed on the front page of the e-LN (Figure 1A). Students can easily add the results of the MUSCLE alignment to their e-LN by using a screen capture (Figure 4A), followed by annotation of the screen image using software such as Google Drawing or YouDoodle and posting as an e-LN entry (Figure 4B). In Figure 4B, a student has indicated the different types of polymorphisms present in the alignment (see also 1). In contrast, the PLN did not typically include bioinformatics data. Instead, these data were often submitted as part of separate, online homework assignment. In our experience, use of the PLN in prior DG courses led students to assign a lower value to bioinformatics data. In contrast, by not artificially separating bench results and sequence analysis data, use of the e-LN helps students understand that bioinformatics data are integral to genome analysis.
After students generate and enter their data into the e-LN during the class period, they complete the e-LN entry at home. Each experiment template includes questions that guide students through data annotation and analysis. Once all of the data have been entered and the analysis is complete, students submit the unpublished entry for review (Figure 4B, red circle). At this point, the grader can review the entry, provide feedback, and assign a grade using Whisper Comments. The grader then publishes the completed notebook entry. The notebook entry is then viewable by all classmates in the fully formatted version (Figure 5), but the Whisper Comments remain private. Thus, students can view and comment on each other's completed entries without seeing any private Whisper Comments. Students can also share draft entries with each other without sharing the Whisper Comments. In this way, students can collaborate to share results during group projects when the notebook entries are in draft form. After an entry is published it is still editable by the student author. In summary, the e-LN (but not the PLN) allows collaborations with all members of the class during and after completion of the experiment while protecting sensitive comments.
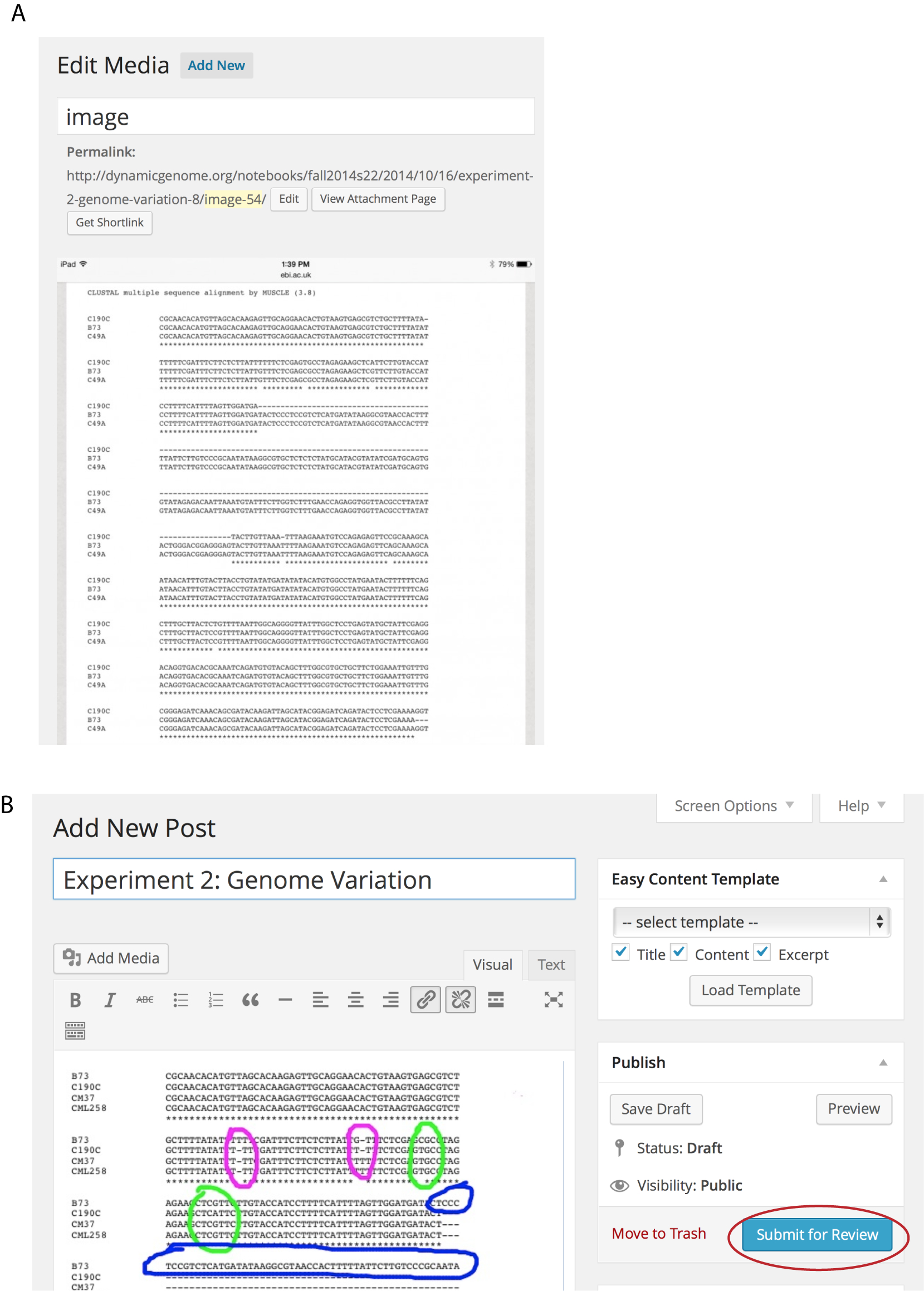
Figure 4. Bioinformatics data loaded into e-LN draft. A. Screen capture of a web page showing multiple sequence alignment output before annotation. B. Same screen capture following annotation. Students use a separate program such as Google Draw to mark up gels and images. The entry is submitted for grading by clicking the “Submit for Review” button (red circle).
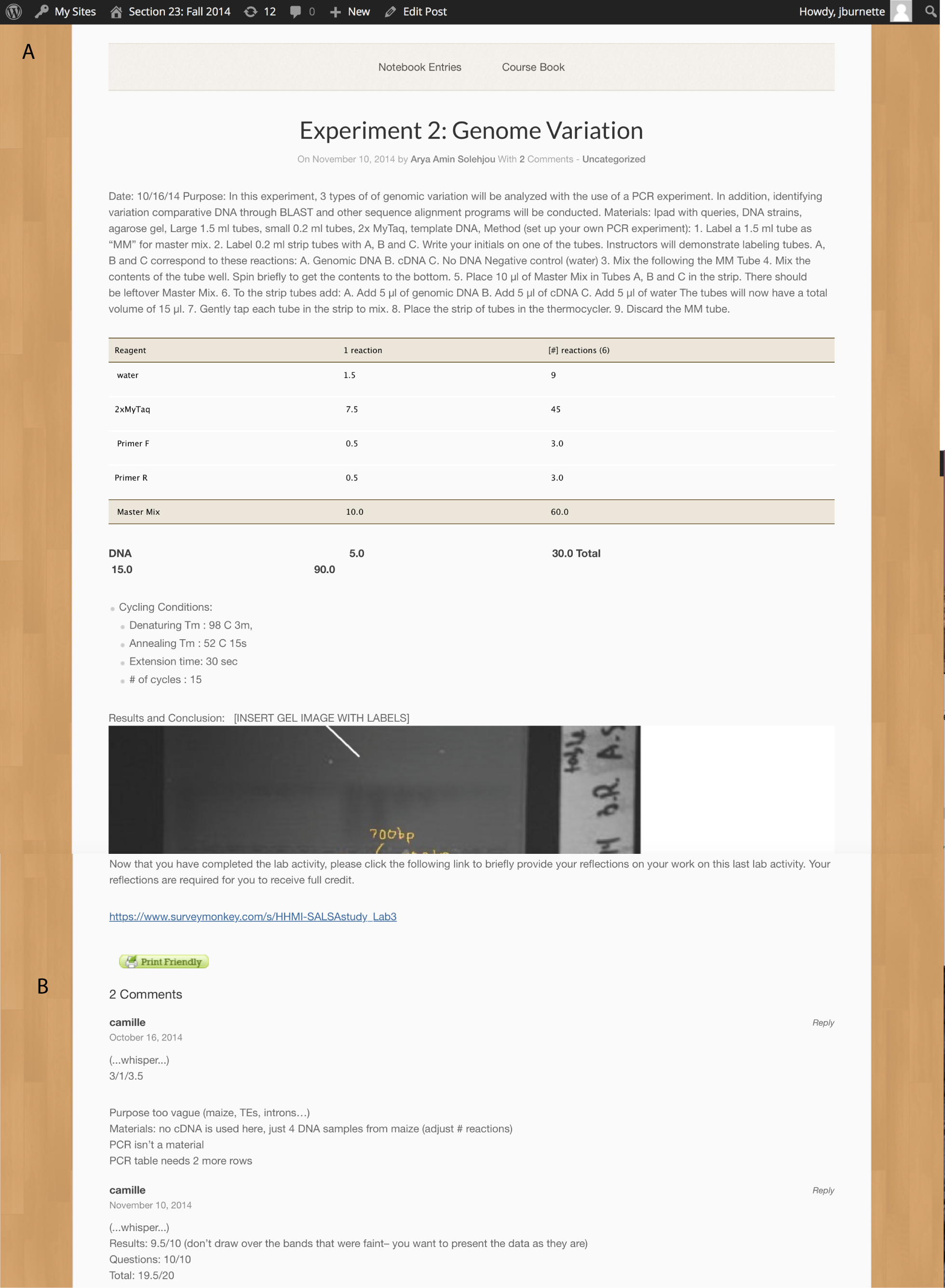
Figure 5. Published post showing Whisper Comments A. The beginning of the Whisper Comment on an e-LN entry. B. The end of the Whisper Comment from the TA. These panels show an actual student entry, complete with grammatical errors. The TA may comment on grammar, but the focus of the grading is on completeness and content. The Whisper Comments appear only on the screens of instructor, the TA, and the student who wrote the notebook entry; they are not visible by other students in the class.
Evaluation of the e-LN
A concern of using technology in the classroom is that it might add an additional burden on the students. To determine how the students perceived the e-LN, we conducted an anonymous survey that consisted of nine questions, that asked students to rank their reaction from strongly disagree to strongly agree. Overall, students responded positively to the e-LN (Figure 6), especially regarding the value of the templates. Further, students found that using the e-LN and the templates better prepared them for using a PLN in other classes or in their research laboratory.

Figure 6. Survey results of opinions about the e-LN. Students were asked to rate their opinion from strongly disagree to strongly agree on nine questions. The average rating is shown.
Inclusivity
Wordpress is compliant with most computer accessibility requirements and thus e-LN is inclusive to all individuals including the sight and hearing impaired (http://en.support.wordpress.com/accessibility/). Taken on its own merit, this accessibility argues strongly for the adoption of an e-LN over a PLN.
CONCLUSION
The e-LN used in our DG course is a free, online documentation system suitable for both the classroom and research laboratory that overcomes many of the limitations of a PLN. A key feature is that it allows instructors to provide timely feedback to students and permits students to collaborate on group projects. The e-LN is not limited to scientific notebooks or record keeping. For example we have included links to surveys for student reflection in the templates. It would also be possible to ask for feedback directly in the template. Lecture only courses could use the e-LN to provide lecture notes and in class two-minute essays or other activities. After class, instructors can review the student entries and give feedback to individual students. In conjunction with Google Drawing students could create figures in class for study use later. Further these data are captured for potential course evaluation purposes. e-LNs are becoming the standard for the documentation of experiments in research laboratories (5). By using the e-LN, students are also preparing for future careers in science and medicine because the Affordable Care Act promotes the use of electronic medical records (6). The majority of our students are "digital natives" and thus comfortable with adopting new technology and software (7). Our students quickly adopted the e-LN and would like to see it deployed in other classes and laboratories.
ACKNOWLEDGMENTS
We thank Dr. Jason Stajich and Biology 20 students and TAs who have used the e-LN and provided helpful feedback to improve the system. This project was supported by a University of California, Riverside Innovative Use of Information Technology (IUIT) grant to S.R.W., an NSF grant (DUE-1317642) to Dr. Michael McKibben and an HHMI Institutional Grant (52008110) to S.R.W. Alexandra Chapovskya and Krista Palmer are undergraduates who helped develop the e-LN templates and tested early versions.
References
- Burnette JM, Wessler SR. 2013. Transposing from the laboratory to the classroom to generate authentic research experiences for undergraduates. Genetics 193:367-375.
- Walsh E, Cho I. 2012. Using Evernote as an electronic lab notebook in ... [J Lab Autom. 2013] - PubMed - NCBI. Journal of Laboratory Automation 18:229-234.
- Hall M, Vardar-Ulu D. 2013. An inquiry-based biochemistry laboratory structure emphasizing competency in the scientific process: A guided approach with an electronic notebook format - L. Hall - 2013 - Biochemistry and Molecular Biology Education - Wiley Online Library. http://onlinelibrary.wiley.com/doi/10.1002/bmb.20769/abstract. Accessed February 4, 2015.
- Johnston J, Kant S, Gysbers V, Hancock D, Denyer G. 2014. Using an ePortfolio system as an electronic laboratory notebook in undergraduate biochemistry and molecular biology practical classes. Biochemistry and Molecular Biology Education 42:50-57.
- Nussbeck SY, Weil P, Menzel J, Marzec B, Lorberg K, Schwappach B. 2014. The laboratory notebook in the 21st century: The electronic laboratory notebook would enhance good scientific practice and increase research productivity. EMBO reports 15:631-634.
- Hsiao C, Hing E. 2012. Use and characteristics of electronic health record systems among office-based physician practices: United States, 2001-2012. NCHS data brief 111.
- Prensky M. 2001. Digital Natives, Digital Immigrants Part 1. On the Horizon 9:1-6.
Article Files
Login to access supporting documents
An Open Source, Collaborative Electronic Notebook for Undergraduate Laboratory Classes(PDF | 2 MB)
022015-0001BTeaching Tools and Strategies-s01.tif(TIF | 12 MB)
- License terms

Comments
Comments
There are no comments on this resource.