Furry with a chance of evolution: Exploring genetic drift with tuco-tucos
Published online:
Abstract
Genetic drift is an important mechanism of evolution, yet undergraduates often fail to understand how it leads to evolutionary change due in part to its random nature. This lesson plan describes a simulation-based activity that allows students to demonstrate the process of genetic drift across generations. Using a simulated population of tuco-tucos - a small rodent native to South America - students can explore how allele frequencies can change over time due to chance. Students will also demonstrate random changes in allele frequency (genetic drift) using two different population sizes (with an extended option for a third population size) so they may better conceptualize the impact of population size on genetic drift as an evolutionary force. Using inexpensive materials (beans and paper cups), instructors can actively engage students in the process of evolution. The simulations are followed by a brief discussion of two real-world examples of bottleneck and founder effects, two events when the impact of genetic drift can become more pronounced. The lesson then ends with a series of thought questions to reinforce student understanding of how genetic drift leads to evolution. This activity is appropriate for small or large class sizes and advanced high school and college biology courses. It can also be adapted for non-major college biology courses.
Citation
Hsu, J., Imad, M., and Wilson, K.M. 2019. Furry with a chance of evolution: Exploring genetic drift with tuco tucos. CourseSource. https://doi.org/10.24918/cs.2019.17Lesson Learning Goals
- Students will understand genetic drift as an evolutionary force and its relationship to randomness.
- Students will understand how population size affects the relative influence of genetic drift.
- Students will understand how bottlenecks and founder effects can lead to an increased impact of genetic drift in populations.
Lesson Learning Objectives
- Students will be able to explain how genetic drift leads to allelic changes over generations.
- Students will be able to demonstrate that sampling error can affect every generation, which can result in random changes in allelic frequency.
- Students will be able to explore and evaluate the effect of population size on the strength of genetic drift.
- Students will be able to analyze quantitative data associated with genetic drift.
Article Context
Course
Article Type
Course Level
Bloom's Cognitive Level
Vision and Change Core Competencies
Vision and Change Core Concepts
Class Type
Class Size
Audience
Lesson Length
Pedagogical Approaches
Principles of How People Learn
Assessment Type
INTRODUCTION
Genetic drift - the change in allele frequency due to chance survival of individuals - is one of the major mechanisms of evolution, along with selection, mutation, and migration between populations. Understanding these four major mechanisms of evolution has been widely highlighted as critical to undergraduate biology education and a deep conceptual understanding of evolution (e.g., Biocore Guide [1]), given that these forces drive changes in all species and can cause population-level changes that lead to the formation of new species. In particular, many students incorrectly equate evolution and natural selection and do not recognize that there are other mechanisms of evolution (2). Even if students have learned about drift, many misconceptions often still remain due in part to drift's random nature and because it creates evolutionary change that is not always beneficial (2,3). A number of papers have documented both its fundamental role in understanding evolution as well as the multiple misconceptions that students hold about genetic drift, including interpreting drift as adaptation to the environment (2-5). This challenge is exacerbated by the abstract nature of genetic drift, confusion over the role of randomness in drift and other biological processes, and the limited depth that most introductory biology textbooks provide for genetic drift (6-7). Indeed, we have found in teaching our own students that students do not perform as well on assessments of genetic drift relative to other mechanisms of evolution. Performance aggregated over several courses revealed that on average, 58% of students correctly answered assessment questions based on genetic drift compared to 74% for questions based on other mechanisms of evolution. Additionally. others have found that nearly 75% of biology undergraduates harbor misconceptions about drift even after instruction (2). Accordingly, we present here a short simulation-based module designed for students in an introductory undergraduate biology course. The aim of this module is to provide students with strong foundational knowledge for one mechanism of evolution by allowing them to explore the random nature of genetic drift through simulating changes in allele frequency across generations.
While there have been other simulation-based modules developed that cover genetic drift (e.g., 8-12), some include other mechanisms of evolution simultaneously (9,10), engendering confusion, or fail to assess changes in allele frequency across generations (8), which is the true measure of genetic drift acting in a population. Lastly, these simulations are limited in that they are only practical for use in small classes. This module was designed for and tested in both small and large lectures and incorporates a wide range of active learning techniques and formative assessments. Thus, we build upon other foundational versions of simulated genetic drift activities (e.g., 8-12) by presenting a lesson that is versatile, in-depth, and reliant on real scenarios to help students understand genetic drift. In addition, our inquiry-based approach guides students through discovering major concepts while providing them with practice in developing quantitative skills. By challenging students to calculate frequencies and graph their own data, this activity promotes connections between math and science, thus supporting students' ability to use quantitative reasoning, an area that has been highlighted as a core competency to biological literacy in Vision and Change (13).
INTENDED AUDIENCE
This module is designed to be implemented in a post-secondary introductory biology course where evolution is introduced and may also be incorporated into non-majors' biology courses or advanced high school biology courses. It has been used successfully in both small (20 students) and large (90 students) class sizes, in introductory and non-majors' biology courses and in community college, liberal arts and research university settings.
REQUIRED LEARNING TIME
This module will take between 40 minutes and one hour, depending on class size, pace, and length of student discussions. See Table 1 for suggested times.
PRE-REQUISITE STUDENT KNOWLEDGE
This module presumes that students have an understanding of evolution and natural selection from previous modules in the biology course. Instructors may choose to discuss the remaining mechanisms of evolution after this module.
PRE-REQUISITE TEACHER KNOWLEDGE
The instructor should have a strong foundation in and capacity to teach evolution. The instructor should specifically have an understanding of how genetic drift functions in a random manner and how population size influences the relative impact of genetic drift acting to change the allele frequencies of a population over time. The instructor should also be comfortable conducting hands-on activities for her/his given class size. While most current general biology textbooks will provide sufficient overviews, several open source resources are also available online to help both students and instructors develop their pre-requisite knowledge (e.g., 14-16).
SCIENTIFIC TEACHING THEMES
ACTIVE LEARNING
Students will work with peers to answer questions using think-pair-share style discussions and to simulate the change in allele frequency over a number of generations using a container with two different colored beans to represent two alleles of the same gene. After each simulation, students will add their results to the class data set either manually (for small classes) or using clicker voting tools (for larger classes). They will analyze class results in order to explore overall trends in changes to allele frequencies both in groups and as a class. Students will also practice visualizing data graphically by plotting their results from each simulation.
ASSESSMENT
Students will answer questions in class provided on the included slides (Supporting File S1: Furry with a chance of evolution - Small lecture slides and S2: Furry with a chance of evolution - Large lecture slides) as well as from the student handout (Supporting File S3: Exploring genetic drift with tuco-tucos - Student handout) as part of the activity. They will receive feedback for questions included on slides immediately in order to build on concepts relating to genetic drift. Students will also receive feedback in their responses to the corresponding handout (Supporting File S3: Student handout) and correct answers will be made available (Supporting File S4: Exploring genetic drift with tuco-tucos - Student handout instructor version). Students will later be asked to apply concepts of genetic drift during the subsequent exam. They will be provided with a novel genetic drift scenario and asked to predict the outcome. An example exam question has also been provided (Supporting File S5: Sample Genetic Drift Questions - Assessment questions).
INCLUSIVE TEACHING
Students will contribute individual data to build an aggregate data set for the class, which will be used to further the discussion of genetic drift. Engaging students in this manner can help them feel a stronger sense of community. Students will also have a chance to build community as they work through genetic drift simulations with one or two peers. Since genetic drift is based on probability, there is a chance that results of the simulations will not be as expected (e.g., no groups will reach fixation, or all groups will reach fixation for the same bean color), which provides an opportunity for students to explore what happens in science when predictions are not met. As an addition to the end of the class activity, the instructor may ask students to generate their own activity or method for simulating genetic drift.
LESSON PLAN
PRE-CLASS PREPARATION
Before the class, decide how many small groups you will form in your class. We recommend two-three students per group. Then, print handouts and obtain the necessary supplies, as listed below:
- Brown beans and white beans of the same size. If beans are not available, any other small object that comes in two different color variants will be acceptable as long as they are the same size, for example, small candies or pompoms (items that roll easily are not recommended). Trials have shown that even slight differences in object size can lead to biased results, which may hinder student understanding. You will need 8 beans of each color per group in your class.
- Small cups to hold and draw beans from (one per group).
- Copies of the student handout (Supporting File S3: Exploring genetic drift with tuco-tucos -Student handout) or blank sheets for students to record their data.
Before class, prepare one cup per group with at least 16 beans (eight of each color) inside each cup (more is ok too). Finally, plan how you would like to collect data from each group in order to make it visible to the class. For example, for a small class, you may ask student groups to record their data on a piece of paper separate from the student handout (Supporting File S3: Student handout) to hand in. You may then choose to use a whiteboard and draw a table/chart ahead of time to record and display the aggregate data from all groups in during class. Alternatively, for larger classes, you may use online collaborative programs (e.g. Google Docs) or clickers. Students will need access to these class data for a later part of the activity.
CLASS ACTIVITY
Background
In order to give context to the activity, begin by first providing an overview of what you have already covered about mechanisms of evolution, and how the topic of genetic drift fits in. You should emphasize that random genetic drift is one of the four main forces that drive evolution; it is highly recommended that you have covered natural selection before this module. A common misconception among high school and first- and second-year biology students is that natural selection is the only mechanism which gives rise to evolution. It may be worthwhile to survey the students about natural selection and evolution.
Next, you should introduce the study system of tuco-tucos, rodents from South America, and explain that you will be focusing on potential evolution in the trait of the pelage (coat) color in tuco-tucos. Pelage color was chosen as an example of a discrete, easily observed trait that varies in certain species of tuco-tucos and is controlled genetically (17,18). We chose tuco-tucos given that they represent one of the authors' study systems, but you should feel free to change to a different organism and different trait, given that the choice of organism and trait will not impact the lesson. Drift will act on any genetic variation in a finite population, although we recommend choosing an easily-observed discrete trait to promote student comprehension. Present the sample data showing a change in allelic frequency of a given gene over time (Supporting Files S1: Furry with a chance of evolution - Small lecture slides, slide 2 and S2: Furry with a chance of evolution - Large lecture slides, slide 2). You will then ask students to think about whether evolution is occurring, and if they think these changes can be driven by random processes, nonrandom processes, or both. You have lots of options here regarding time, degree of interaction between students, and how you capture responses. We recommend that you ask students to think silently about these questions, and then assess their responses using a clicker system or asking them via a show of hands/fingers. We recommend a think-pair-share strategy along with or in conjunction with the above two response methods. Students will likely provide a mix of responses; we recommend that you not provide the correct responses until the end of this background section but use the discussion to help students think about what it means for a population to be evolving in order to prime them for the topic of genetic drift. Thus, these opening thought questions allow students to make predictions and apply concepts they have learned in previous classes to a new situation.
Opening activity - genetic drift simulation
After these opening thought questions, you should then introduce the activity, where each group will simulate genetic drift in tuco-tucos using beans of two different color to represent variation in tuco-tuco pelage (coat) color, a trait with a genetic basis. For the sake of simplicity, this activity assumes haploidy so that a single bean may represent a single tuco-tuco. Be sure to explain that for the purposes of this simulation this is not a trait that is being acted upon by natural selection (i.e. a tuco-tuco is equally likely to survive no matter its coat color). We recommend first explaining the steps, and then demonstrating the simulation for two generations using a cup of beans just like the students have. Consider also explaining how simulations work, in general, and how computational biologists use modeling and simulations to inform our understanding of evolution. Be sure to explain each step again as you demonstrate it.
1. Start with four beans of each color (for a total of eight beans). Set aside all additional beans for now. These eight beans represent your population of eight tuco-tucos.
2. Without looking, randomly select four beans to represent the individuals that will contribute their genes to the next generation. You can have any of the following combinations: four white, zero brown; three white, one brown; two white, two brown; one white, three brown; zero white, four brown.
3. Simulate reproduction by duplicating the selected four beans so that you again have eight beans. The new population should have the same ratio of white to brown as the four beans selected in step two. (This is done in order to maintain the same population size of eight.) This means that if your random selection from step two yielded one white and three brown you will have two white and six brown by the end of this step.
4. Record the number of brown and white beans next to the generation number starting with 'generation 1' (Supporting File S3: Student handout, Part I).
5. Repeat steps two-four for five minutes or until all beans are a single color (eight white or eight brown). Each new random selection represents a new generation and should be recorded next to the subsequent generation (Supporting File S3: Student handout, Part I).
As a demonstration round:
- You begin by placing your population of four white and four brown beans on your desk.
- You look away and randomly select three white beans and one brown bean.
- You will then double this to have six white and two brown beans. At the end of this step, have the students pause and ask them what the numbers mean. Remind them that this will represent a random sample of their population surviving and reproducing to generate the next generation.
- For 'generation 1' (Supporting File S3: Student handout, Part I), you will record '6' in the column for white beans and '2' in the column for brown beans.
- From this new population, you will randomly select four beans again and repeat the process recording your results for the subsequent generation.
Following this demonstration, instruct students to follow this same process and repeat for five minutes or until they have beans of all the same color. If students finish early, you may instruct them to work on the first two questions for Part I (Supporting File S3: Student handout, Part I) or ask them to run a second simulation to add to the class data. You should then have students record their data (using your pre-determined method of data collection, either on the board, using a projected online document or clicker polling) of 1) the final number of white and brown colored tuco-tucos in their populations and 2) the number of generations they reached.
Allow students a few minutes to discuss the first two questions for Supporting File S3: Student handout, Part I (also in Supporting File S1: Small lecture slides, slide 10 and Supporting File S2: Large lecture slides, slide 11) and observe any trends that stand out in the class data. Use the class results to guide your discussion on how genetic drift can lead to evolution and to answer the first two questions. For example, highlight that some groups ended at 50:50 white to brown while others ended at 0:100 white to brown colored tuco-tucos. Introduce and define fixation, extinction of trait variants (or alleles), and genetic drift. Also discuss if there was any pattern in: 1) which color bean reached fixation, and 2) the number of generations it took to reach fixation. Most classes should have close to an even divide in the number of groups reaching fixation for each color, but there will likely be variance in how quickly it takes each group to reach fixation. Highlight this observation and ask students to think about why this is, and how this is relevant to evolution. To wrap up this part of the activity, you should emphasize how the students have now simulated a random process that can lead to evolution, and that natural selection is thus not the only mechanism of evolution.
Part II of activity - an exploration of population size on genetic drift
In the second part of the class activity, you will lead students through a guided inquiry simulation where students critically think about how population size could affect the strength of genetic drift that was just modeled. You should first ask them to make predictions on how changing population size (i.e., the number of tuco-tucos) would impact 1) the probability of any one trait reaching fixation and 2) the time it takes to reach fixation. Then ask students to redo the simulation but with half as many beans (two white and two brown), and again gather data from the class. This second simulation should go very quickly since most groups will reach fixation within a few generations. Following the data collection, ask students to identify and explain the differences between the two simulations, and provide the additional thought questions (Supporting File S1: Small lecture slides, slide 12, S2: Large lecture slides, slide 14 and S3: Student handout, Part II) on simulations with smaller or larger population sizes. We recommend that you use a combination of think-pair-share and either the clicker system or show of hands to guide your discussion. During your discussion, you should formally define the relationship between genetic drift and population size, i.e., the larger the population size the longer it will likely take to reach fixation due to drift. If time permits, you can expand on this by simulating the effect of genetic drift on a larger population by increasing the population size to 16 tuco-tucos. We recommend you set a time limit or generation limit for this activity since it could take much longer than the others. If you don't have time for this part during class, you may ask the students to do it at home.
Part III of activity - founder effect, bottleneck, and wrap-up
In the final part of the activity, you will have students think about what real-life scenarios create small populations, thus increasing the impact of genetic drift. You may choose to do a think-pair-share for this or a more general class discussion only. You should guide the discussion to cover bottlenecks and founder effects, both of which result in populations with reduced size and thus show a greater impact of genetic drift. Real-life scenarios for both bottlenecks and founder effects are included in Supporting File S1: Small lecture slides, slides 13-16 and S2: Large lecture slides, slides 15-18 as examples. Ask the students to reflect on why it is important to understand genetic drift and how it might impact them directly. Point out the differences between natural selection and genetic drift. Natural selection leads to the propagation of traits beneficial for survival and reproduction. Since genetic drift is random, it can lead to an increase in the frequency of traits that are potentially harmful or neutral to fitness. Note that it is still possible for genetic drift to cause an increase in the frequency of beneficial traits, though.
Following this, you should provide the take-home points for this module, then return to the opening two thought questions and ask students to re-submit their answers. You can use these as a formative assessment, where students should now identify that the population is evolving and that the change can be due to both random and non-random processes, where both drift and selection can act simultaneously on a population. You may also introduce the third thought question here, where students think about how their answers may change if the examined gene is known to not be under any selective pressure. Students should be able to now say that the changes in allele frequency are due solely to drift (assuming a closed population with no mutations). This third question will help students compare genetic drift to natural selection in order to reinforce that genetic drift is a random process and understand that more than one evolutionary process may act on a single trait simultaneously.
Lastly, Supporting File S3: Student handout contains space for students to graph the results of their simulations. Graphing may be done as part of the activity during parts I and II, homework, additional practice, or omitted. The aim of the graphing activity is to help students visualize their results in order to strengthen their understanding of the patterns generated by their data. The graphing activity is also a great way to help students practice making and interpreting figures.
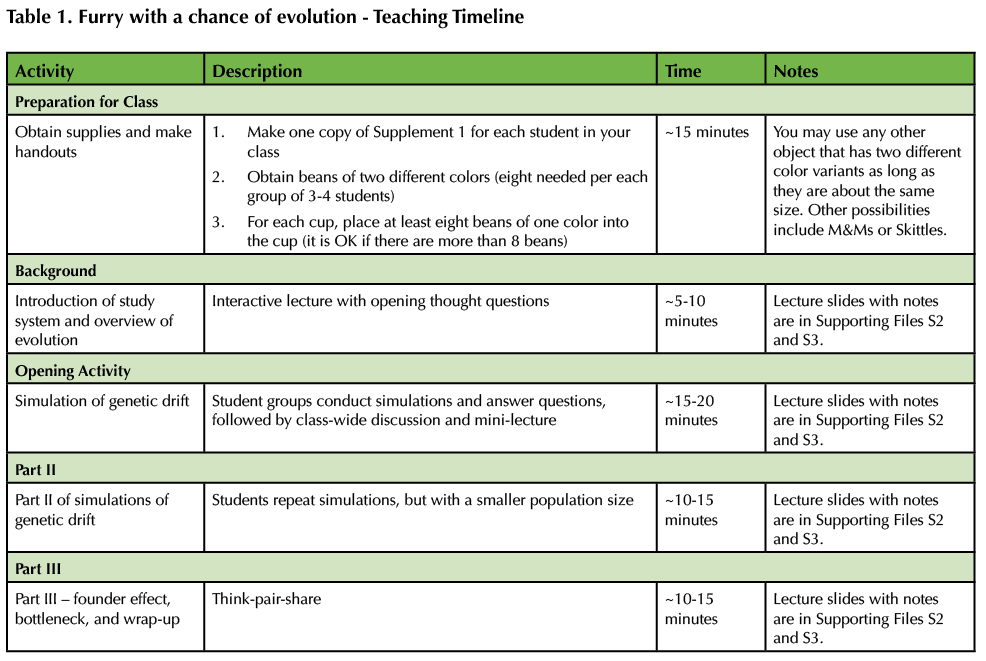
Table 1. Furry with a chance of evolution - Teaching Timeline
TEACHING DISCUSSION
The lesson was very well received by students and was beneficial in helping students overcome common misconceptions surrounding genetic drift. Student evaluations and unsolicited comments indicated that students enjoyed using the simulation as a strategy to learn about evolution. Compared to responses to thought questions at the beginning of the activity (Supporting Files S1: Small lecture slides, slide 2 and S2: Large lecture slides slide 2), student understanding of genetic drift improved after the activity when the same questions plus an additional one were asked (Supporting Files S1: Small lecture slides, slide 18 and Supporting File S2: Large lecture slides, slide 20). Additional in-class formative assessments following a larger lesson on evolution showed that over 90% of students were able to identify genetic drift as a non-random process and an evolutionary force that could result in a reduction in genetic variation. In addition, instructors may wish to discuss scientific studies relevant to the chosen study organism used in this activity. For instance, if instructors kept in the choice of the tuco-tuco, then instructors can discuss how in certain species of tuco-tucos there is natural variation in pelage (fur) color, where some are melanic (dark-colored), agouti (alternating bands of dark and light-colored pigmentation), or dark-backed (17-18). While this trait is likely under selective pressure in the wild, several studies have examined the role of genetic drift in driving shifting pelage color frequencies in various populations of tuco-tucos (17-18). In addition, other species of tuco-tucos have undergone natural bottlenecks triggered by rapid environmental changes (e.g., floods and volcanic eruptions) (19-23). These natural events that increase the impact of drift through the reduction of population size thus offer instructors a unique opportunity to connect to recent events and scientific studies.
For larger lectures, we recommend that you use slides from Supporting File S2: Large lecture slides in place of the slides from Supporting File S1: Small lecture slides. Supporting File S2: Large lecture slides contain clicker voting questions to make data collection after each round faster and will allow you to display the results graphically. When you demonstrate the activity, ask for questions after your initial demonstration. If you are concerned that students don't know how to proceed, we recommend that you repeat the demonstration a second time. Even in large classes, it will be important to facilitate the activity by walking around to individual groups. Lastly, as an alternative to asking students to write their answers to questions on the included worksheet (Supporting File S3: Student handout), you may instead: 1) ask them to write their answers as a discussion post on your learning management system (e.g. Blackboard, Canvas, Moodle, D2L, etc.), 2) discuss the questions in a think-pair-share format, or 3) use the questions as a 1-minute paper prompt. It will be important to implement a strategy that will allow you to review responses to identify any remaining misconceptions.
This activity was also piloted in a non-majors general education biology course. The course focused on topics such as genetics, biotechnology, and human affairs. The post-activity assessment indicated that it was effective at improving student understanding of genetic drift. However, if instructors wish to use this activity for non-majors, we recommend they leave extra time (as much as 100 minutes) to introduce background information and complete the activity. It is imperative to establish a baseline understanding of underlying concepts like the relationship between a trait and an allele, and that mechanisms of evolution other than natural selection exist. Other knowledge gaps will exist that will be specific to the course implementation. We also recommend taking more time to explore examples where genetic drift becomes more pronounced (bottleneck and founder effect) so students can better understand how genetic drift may act and how these can lead to poorly adapted populations.
SUPPORTING MATERIALS
- S1. Furry with a chance of evolution - Small lecture slides. Presentation slides to be used when teaching the lesson. These slides are recommended for smaller class sizes but may also be used for larger classes.
- S2. Furry with a chance of evolution - Large lecture slides. Alternative presentation slides to be used when teaching the lesson. These slides contain clicker polls to collect data on allele frequencies and generation times and are recommended for larger class sizes but may also be used for smaller classes.
- S3. Furry with a chance of evolution - Student handout. Accompanying worksheet to be used concurrently with the activity. A portion of the worksheet (Extension Questions) is designed to be completed at home, but we encourage instructors to allow class time for the entire worksheet if time allows.
- S4. Furry with a chance of evolution - Student handout instructor version. The answers provided here will help instructors assess student responses.
- S5. Furry with a chance of evolution - Sample assessment questions. This includes a multiple-choice exam question and references for the Genetic Drift Concept Inventory and related work examining misconceptions about drift.
ACKNOWLEDGMENTS
We would like to thank Keefe Reuther and Brian Dalton for their contributions to the supplementary materials and input on teaching the module and Nancy Burley for her comments on an earlier version of this manuscript. We would also like to thank Kevin Wilson for photographic contributions to the supporting materials.
References
- Brownell SE, Freeman S, Wenderoth MP, Crowe AJ. 2014. BioCore guide: A tool for interpreting the core concepts of Vision and Change for biology majors. CBE-Life Sciences Education. 13: 200-211.
- Andrews TM, Price RM, Mead LS, McElhinny TL, Thanukos A, Perez KE, Herreid CF, Terry DR, Lemons PP. 2012. Biology undergraduates' misconceptions about genetic drift. CBE-Life Sciences Education. 11:248-259.
- Summers MM, Couch BA, Knight JK, Brownell SE, Crowe JA, Semsar K, Wright CD, Smith MK. 2018. EcoEvo-MAPS: An ecology and evolution assessment for introductory through advanced undergraduates. CBE-Life Sciences Education 17: ar18, 1-12.
- Price RM, Andrews TC, McElhinny TL, Mead LS, Abraham JK, Thanukos A, Perez KE. 2014. The genetic drift inventory: A tool for measuring what advanced undergraduates have mastered about genetic drift. CBE-Life Sciences Education. 13:65-75.
- Klymkowsky M, Garvin-Doxas K. 2008. Recognizing student misconceptions through Ed's Tools and the Biology Concept Inventory. PLoS Biol. 6:14-17.
- Garvin-Doxas K, Klymkowsky M. 2008. Understanding randomness and its impact on student learning: Lessons learned from building the Biology Concept Inventory (BCI). CBE-Life Sciences Education. 7:227-233.
- Beggrow EP, Nehm RH. 2012. Students' mental models of evolutionary causation: Natural selection and genetic drift. Evolution (N Y). 5:429-444.
- Maret TJ, Rissing SW. 1998. Exploring genetic drift and natural selection through a simulation activity. The American Biology Teacher. 60: 681-683.
- Staub NL. 2002. Teaching evolutionary mechanisms: Genetic drift and M&M's. Bioscience 52: 373-377.
- Russo CAM, Voloch CM. 2012. Beads and dice in a genetic drift exercise. Evolution: Education and Outreach. 5: 494-500.
- Mukhopadhyay CC, Henze R, and Moses YT. 2014. If Not Race, How Do We Explain Biological Differences?" in: How Real is Race? A Sourcebook on Race, Culture, and Biology. 2nd Edition. 2014. AltaMira Press.
- Genetic Drift Exercise. 2005. University of Virginia Biology Department. Available online at http://www.faculty.virginia.edu/evolutionlabs/genetic_drift_exercise.html
- AAAS (2011). Vision and Change in Undergraduate Biology Education: A Call to Action. Washington, DC: AAAS. Available online at http://visionandchange.org/finalreport/.
- Understanding Evolution. 2019. University of California Museum of Paleontology. Available online at http://evolution.berkeley.edu/.
- Kliman R, Sheehy B, Schultz J. 2008. Genetic drift and effective population size. Nature Education 1:3.
- Kimura M, Ohta T. 1969. The average number of generations until fixation of a mutant gene in a population. Genetics 61, 763-771.
- Wlasiuk G, Garza JC, Lessa EP. 2003. Genetic and geographic differentiation in the Rio Nego tuco-tuco (Ctenomys rionegrensis): Inferring the roles of migration and drift from multiple genetic markers. Evolution 57: 913-926.
- D'Anatro A, Lessa EP. 2006. Geometric morphometric analyses of geographic variation in the Rio Negro tuco-tuco, Ctenomys rionegrensis (Rodentia: Ctenomyidae). Mammalian Biology 71: 288-298.
- Chan YL, Anderson CN, Hadly EA. 2006. Bayesian estimation of the timing and severity of a population bottleneck from ancient DNA. PLoS Genetics 2: e59.
- Fernandez-Stolz GP, Stolz JFB, de Freitas TRO. 2007. Bottlenecks and dispersal in the tuco-tuco das dunas, Ctenomys flamarioni (Rodentia: Ctenomyidae), in Southern Brazil. J Mammalogy 88: 935-945.
- Kittlein MJ, Gaggiotti OE. 2008. Interactions between environmental factors can hide isolation by distance patterns: A case study of Ctenomys rionegrensis in Uruguay. Proc R Soc B 275: 2633-2638.
- Hsu JL, Kam S, Tammone MN, Lacey EA, Hadly EA. 2017. Rapid increase in genetic diversity in an endemic Patagonian tuco-tuco following a recent volcanic eruption. J Mammalogy 98: 779-792.
- Hsu JL, Crawford JC, Tammone MN, Ramakrishnan U, Lacey EA, Hadly EA. 2017. Genomic data reveal a loss of diversity in two species of tuco-tucos (genus Ctenomys) following a volcanic eruption. Scientific Reports 7: 16227.
Article Files
Login to access supporting documents
Furry with a chance of evolution: Exploring genetic drift with tuco-tucos(PDF | 126 KB)
S1. Furry with a chance of evolution - Small lecture slides.pptx(PPTX | 5 MB)
S2. Furry with a chance of evolution - Large lecture slides.pptx(PPTX | 5 MB)
S3. Furry with a chance of evolution - Student handout.docx(DOCX | 300 KB)
S4. Furry with a chance of evolution - Student handout instructor version.docx(DOCX | 300 KB)
S5. Furry with a chance of evolution - Sample assessment questions.docx(DOCX | 15 KB)
- License terms

Comments
Comments
There are no comments on this resource.