The Impact of Diet and Antibiotics on the Gut Microbiome: Distance Education Variant
Editor: Kristin Fox
Published online:
Abstract
The goal of this article is to describe a variation of an active learning exercise that was previously published by the same author under a similar title. The variation describes modifications instructors can use to make the exercise suitable for online course delivery. The exercise is split into several parts. Part I is taught asynchronously via three consecutive videos. Part II is taught synchronously via Blackboard Collaborate Ultra (or similar). There is a follow up assignment that students do in groups as part III. The active learning exercise is a 'pasta' simulation of the gut microbiome. In the asynchronous part I of this exercise, students are virtually given a plastic bag/gut with different types of pasta/gut bacteria. Six different bags resemble the gut microbiome under six different diets. The instructor mimics an antibiotic treatment by removing two types of pasta/gut bacteria and replacing them with beans/environmental bacteria from a second plastic bag. In the synchronous part II of the exercise, students read multiple review articles and assign bacterial names to the pasta types under the respective diet. They then use the same articles to identify metabolic byproducts that these bacteria produce. In a follow up assignment that constitutes part III, students investigate signal transduction pathways in the human host cells and the potential diseases that can result from a high fat diet.
Original lesson: The Impact of Diet and Antibiotics on the Gut Microbiome
Citation
Pruß BM.2021. The impact of diet and antibiotics on the gut microbiome: Distance education variant. CourseSource. https://doi.org/10.24918/cs.2021.25Society Learning Goals
Microbiology
- Metabolism
- How does the survival and growth of any microorganism in a given environment depend on its metabolic characteristics?
- Systems
- How do microorganisms, cellular and viral, interact with both human and non-human hosts in beneficial, neutral, or detrimental ways?
- How do microorganisms interact with their environment and modify each other?
Article Context
Course
Article Type
Course Level
Bloom's Cognitive Level
Vision and Change Core Competencies
Vision and Change Core Concepts
Class Type
Class Size
Audience
Lesson Length
Pedagogical Approaches
Principles of How People Learn
Assessment Type
INTRODUCTION
The human microbiome is the compilation of all bacteria, fungi, and sometimes viruses that live and metabolize in and on our bodies and contribute to our health. The composition of this microbiome changes with age and environmental factors, including diet and antibiotic treatments. Such changes start a signal cascade involving the microbes and the human host that can ultimate lead to chronic inflammation (1). The active learning exercise (guided by Supporting Files S1, S2, and S3 Diet and Gut Microbiome – Videos 1, 2, and 3 for asynchronous instruction) leads the students down a signal transduction pathway that starts with changes in the composition of the gut microbiome in response to a change in diet. The new bacteria secrete metabolic byproducts into the host intestine, further leading to molecular changes in the intestinal cells. Ultimately, this causes a range of inflammatory diseases. As an example of a different type of environmental change, Video 3 (S3. Diet and Gut Microbiome – Video 3 for asynchronous instruction) includes a simulation of an antibiotic treatment that leads to infection with Clostridium difficile for some of the student groups. C. difficile is a common infection that predominantly occurs after antibiotic treatment (2), when a broad-spectrum antibiotic clears the intestine of some, but not all bacteria (3).
The pasta simulation was originally described by Anne Estes and developed for an undergraduate General Microbiology class (4). The author of this manuscript adjusted the exercise to suit an upper-level Bacterial Physiology course and included a detailed analysis of the gut microbiome, as well as the signal transduction cascade that is initiated in response to diet (5). The advanced aspect of the exercise was facilitated by several review articles that were provided to the students (6,7), together with information on the Human Microbiome Project (8). To allow for online delivery of this exercise, this article describes an asynchronous part I of the pasta simulation by Estes (4) and a synchronous part II of the advanced exercise by Prüβ (5). The follow up assignment that constitutes part III has not changed from the previous manuscript (5). Note that learning objectives and assessments are unchanged from the previously published exercise (5), detailed references are also provided in this original manuscript (5).
PART I: ASYNCHRONOUS PASTA SIMULATION
Following the instructions in Table 1, the author filled six plastic bags with pasta and produced three consecutive videos (Supporting Files S1, S2, and S3 Diet and Gut Microbiome – Videos 1, 2, and 3 for asynchronous instruction). The videos can be provided to the students via Blackboard or other learning management system, by email, or as a shared document (e.g., using Google). The videos take a total of approximately 20 min to watch. The instructor is advised to let students know they need a pen and notebook for Video 2 (Supporting File S2). S1. Diet and Gut Microbiome – Video 1 for asynchronous instruction explains the exercise and introduces the students to the six different pasta bags and the six bean bags. In S2. Diet and Gut Microbiome – Video 2 for asynchronous instruction, the author opens the bags and gives the students an opportunity to get a better view at the content. Students are expected to write down the types of pasta that are in each bag, using the descriptive language for the pasta types that is used in Table 1 and the videos. S3 Diet and Gut Microbiome – Video 3 for asynchronous instruction is the antibiotic treatment. The author removes two types of pasta. The author then fills the bags up with environmental bacteria from the bean bags. All pasta bags now contain some of the original pasta and the new beans. Some of the pasta bags also contain lentils. Lentils simulate Clostridium difficile. Yellow lentils simulate a non-toxin producing strain, green lentils a toxin producing variety.
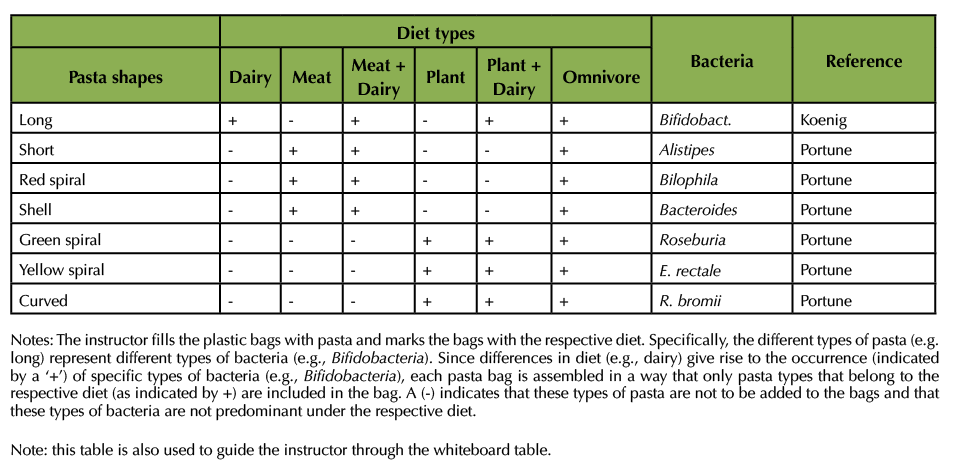
Table 1. Content of the pasta bags.
PART II: SYNCHRONOUS CLASSROOM EXERCISE
This part of the exercise is very similar to the original exercise (5) that was described by this author. Instructors use Blackboard Collaborate Ultra or some other appropriate learning management system (or Zoom or Microsoft Teams). The exercise depends on using a digital whiteboard. Using the whiteboard, the instructor constructs an outline of Table 1, including the different diets and pasta types but without the pluses and minuses, the bacterial names, and the references.
Task 1:
Students report the findings from S2. Diet and Gut Microbiome – Video 2 for asynchronous instruction (5 min). The instructor records on the whiteboard table which pasta types were included in the dairy diet bag, the meat diet bag, etc.
Task 2:
Students use the Nature paper on the Human Microbiome Project (8) to determine the nine phyla of bacteria that contribute to the gut microbiome (5 min). These are Firmicutes, Bacteriodetes, Actinobacteria, Proteobacteria, Fusobacteria, Tenericutes, Spirochaetes, Cyanobacteria, and Verrucomicrobia. Students report their findings to the instructor.
Task 3:
Students use the Koenig paper (6) to determine the predominant bacteria in the gut microbiome of a person who lives on a dairy diet (5 min). Among the first bacteria to colonize the near-sterile gut of a newborn baby, Bifidobacteria are especially important. These are provided to the baby by the mother's milk and aid the degradation of oligosaccharides in the milk. Students report Bifidobacteria as an indicator of a dairy diet and the instructor include this information in the whiteboard Table.
Task 4:
Students use Table 1 of the Portune paper (7) to determine some of the predominant bacteria in adult humans who live on a diet that consists predominantly of meat (5 min). Examples of these bacteria are Alistipes, Bilophila, and Bacteriodes. Students find these in Table 1 and the instructor adds the bacteria to the whiteboard table.
Task 5:
Students use the same Table 1 to determine some of the predominant bacteria in adult humans who live on a diet that consists predominantly of plants (5 min). Examples of these bacteria are Roseburia, Eubacterium rectale, and Ruminoccus bromii. Students find these in Table 1 and the instructor adds the bacteria to the whiteboard table.
Tasks 6 to 8 (5 min):
Repeat tasks 3 to 5 for meat and dairy diet, plant and dairy diet, and omnivore diet. There should now be 20 min left of class time.
Tasks 9 to 11 (20 min):
Students use Tables 1, 2, and 3 of the Portune paper (7) to identify bacteria that either increase or decrease in number following a change in diet to high fat, high protein, and high fiber. If time permits, they identify metabolic byproducts that some of these bacteria secrete into the lumen of the gut. As one example, high fiber diet leads to an increase in Bifidobacteria and Lactobacilli. These are considered probiotics and produce short chain fatty acids including acetate, propionate, and butyrate (9). This combination lowers cholesterol and triglycerides (10). Students report their findings to the instructor. The in-class session is concluded at this time.
PART III: FOLLOW UP ASSIGNMENT
This part of the activity is identical to the original paper by the same author (5). Briefly, students get together in groups of three and study the effect of a change to high fat. They follow the signal transduction cascade, starting with the change in bacteria, the secreted metabolic byproducts, the molecular changes in the host cells, and the ultimate disease. Students are expected to start with the Portune paper (7) and follow additional papers that are referenced in this paper. The assignment is expected to be about two pages of text plus a figure and references. Specific assignment instructions should be given to the students and are provided in the original manuscript, The Impact of Diet and Antibiotics on the Gut Microbiome, by this author (5) as Supporting File S1. Impact of diet and antibiotics – Assignment instructions that instructors give to students for the follow up assignment. A sample assignment is provided in the same lesson as Supporting File S2. Impact of diet and antibiotics – Sample follow up assignment for instructors (5), as well as the grading rubric in Supporting File S3. Impact of diet and antibiotics – Grading rubric for the follow up assignment (5).
An intriguing extension of the study, particularly for an upper level course or a class of pharmacy students could be to have students compare the effect of a broad spectrum and a narrow spectrum antibiotic. Which one is more likely to lead to C. difficile infection?
SUPPORTING MATERIALS
- S1. Diet and Gut Microbiome – Video 1 for asynchronous instruction (https://youtu.be/XVTkdjWW8Uk)
- S2. Diet and Gut Microbiome – Video 2 for asynchronous instruction (https://youtu.be/1au0PUXwZww)
- S3. Diet and Gut Microbiome – Video 3 for asynchronous instruction (https://youtu.be/zGNPK0Fi0xU)
ACKNOWLEDGMENTS
The teaching portion of the author's appointment is funded by the College of Agriculture, Food Systems, and Natural Resources at North Dakota State University. The author participated in an active learning workshop that was funded by NSF grant 1525056.
References
- Ercolini D, Fogliano V. 2018. Food design to feed the human gut Microbiota. J. Agric. Food Chem. 66:3754-58. doi:10.1021/acs.jafc.8b00456.
- Czepiel J, Dróżdż M, Pituch H, Kuijper EJ, Perucki W, Mielimonka A, Goldman S, Wultańska D, Garlicki A, Biesiada G. 2019. Clostridium difficile infection: review. Eur. J. Clin. Microbiol. Infect. Dis. 38:1211-1221. doi:10.1007/s10096-019-03539-6.
- Iizumi T, Battaglia T, Ruiz V, Perez Perez GI. 2017. Gut microbiome and antibiotics. Arch. Med. Res. 48:727-723. doi: 10.1016/j.arcmed.2017.11.004
- Estes, AM. 2015. Modeling the dynamic digestive system microbiome. J. Microbiol. Biol. Educ. 16:271-73. doi:10.1128/jmbe.v16i2.908.
- Prüβ BM. 2019. The impact of diet and antibiotics on the gut microbiome. CourseSource doi:10.24918/cs.2019.28.
- Koenig JE, et al. 2011. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. U S A. 108: 4578-85. doi:10.1073/pnas.1000081107.
- Portune KJ, et al. 2017. Gut microbiota, diet, and obesity-related disorders-the good, the bad, and the future challenges. Mol. Nutr. Food Res. 61.1: 1600252. doi:10.1002/mnfr.201600252.
- Human Microbiome Project Consortium. 2012. Structure, function and diversity of the healthy human microbiome. Nature 486(740):207-214. doi:10.1038/nature11234.
- Costabile A, Klinder A, Fava F, Napolitano A, Fogliano V, Leonard C, Gibson GR, Tuohy KM. 2008. Whole-grain wheat breakfast cereal has a prebiotic effect on the human gut microbiota: a double-blind, placebo-controlled, crossover study. Br. J. Nutr. 99:110-120. doi:10.1017/S0007114507793923.
- Wright RS, Anderson JW, Bridges SR. 1990. Propionate inhibits hepatocyte lipid synthesis. Proc. Soc. Exp. Biol. Med. 195:26-29. doi:10.3181/00379727-195-43113.
Article Files
Login to access supporting documents

Comments
Comments
There are no comments on this resource.