Workshop Report: Summer 2020 Virtual CRISPR in the Classroom
Editor: William Morgan
Published online:
Abstract
As innovations and developments in genome editing technologies using CRISPR-Cas systems progress, the need to disseminate relevant knowledge and build skills among the next generation of young scientists in undergraduate classrooms is vital. Our efforts to enable undergraduate educators to bring CRISPR into their classrooms through in-person workshop training began in 2017 and went virtual during summer of 2020 under COVID-19 lockdown. In this report, we describe the proceedings of the virtual workshop and the feedback we received from the participants. An overwhelming majority of attendees reported that the virtual workshop facilitated gains in learning about CRISPR biology and experimental design. The plans shared by attendees to incorporate both virtual and hands-on CRISPR resources into their courses highlights the impact of this virtual CRISPR in the Classroom Workshop on educator confidence, and the likelihood of attendees to add CRISPR biology to their curriculum after participating in such a workshop.
Primary image: A summary diagram of a virtual workshop on bringing CRISPR biology into the classroom. The molecular rendering created with Illustrate software features Cas9 (gray) in complex with guide RNA (red) and target DNA (blue) (PDB ID: 4OO8, (1, 2)).
Citation
Kubera C, Nivillac NMI, Tanner S, Le P, Begde D, Wolyniak MJ, Challa AK. 2022. Workshop Report: Virtual CRISPR in the Classroom. CourseSource. https://doi.org/10.24918/cs.2022.17Article Context
Course
Article Type
Course Level
Bloom's Cognitive Level
Vision and Change Core Competencies
Vision and Change Core Concepts
Class Type
Audience
Pedagogical Approaches
Principles of How People Learn
Background
CRISPR technology has revolutionized molecular biology research within the last decade. It opened exciting new frontiers for exploring how our genes work together to sustain life and also made unprecedented headway into treating human genetic disorders (e.g., sickle cell anemia) and modifying plant and animal livestock for improved productivity. Harnessing this naturally occurring bacterial adaptive immunity mechanism for gene editing has become an important tool for molecular biology in a relatively short time and an essential skill for trainees who wish to join the research community. The importance of CRISPR technology was confirmed by the award of the 2020 Nobel Prize in Chemistry to the scientists who contributed to the broad use of this technology for gene editing (3). Undergraduates interested in the life sciences and medicine should therefore become proficient with how CRISPR technology works and how it may be used. However, many undergraduate instructors lack the exposure, background knowledge, practical skills and therefore confidence to provide this training to students.
To help provide the training necessary for undergraduate instructors to teach about CRISPR-Cas systems and their applications in research, we offered multi-day workshops for undergraduate instructors from diverse institutions in 2018 and 2019 supported by the National Science Foundation (Award Numbers 1823595 and 1916486 with workshops conducted at The Ohio State University (OSU) and The University of Minnesota-Twin Cities (UMN), respectively) (4). These were expanded from a workshop first offered in 2017 as part of the American Biology Laboratory Educators annual meeting. At these workshops, trainees received instruction on how to use a laboratory workflow designed with undergraduate students in mind and to guide a class through the design and implementation of a CRISPR based research project utilizing zebrafish (Danio rerio). Feedback from the first hands-on workshop (at OSU) allowed us to expand our offerings in the second workshop (at UMN) to include experts proficient in using CRISPR-Cas systems in a variety of additional model systems, including bacteria (Escherichia coli), roundworm (Caenorhabditis elegans), and plants (Arabidopsis thaliana). Participants also spent a portion of the workshop planning how they would implement CRISPR-Cas modules in their own classrooms in an accessible and engaging manner.
CRISPR in the Classroom 2020 Virtual Workshop Report
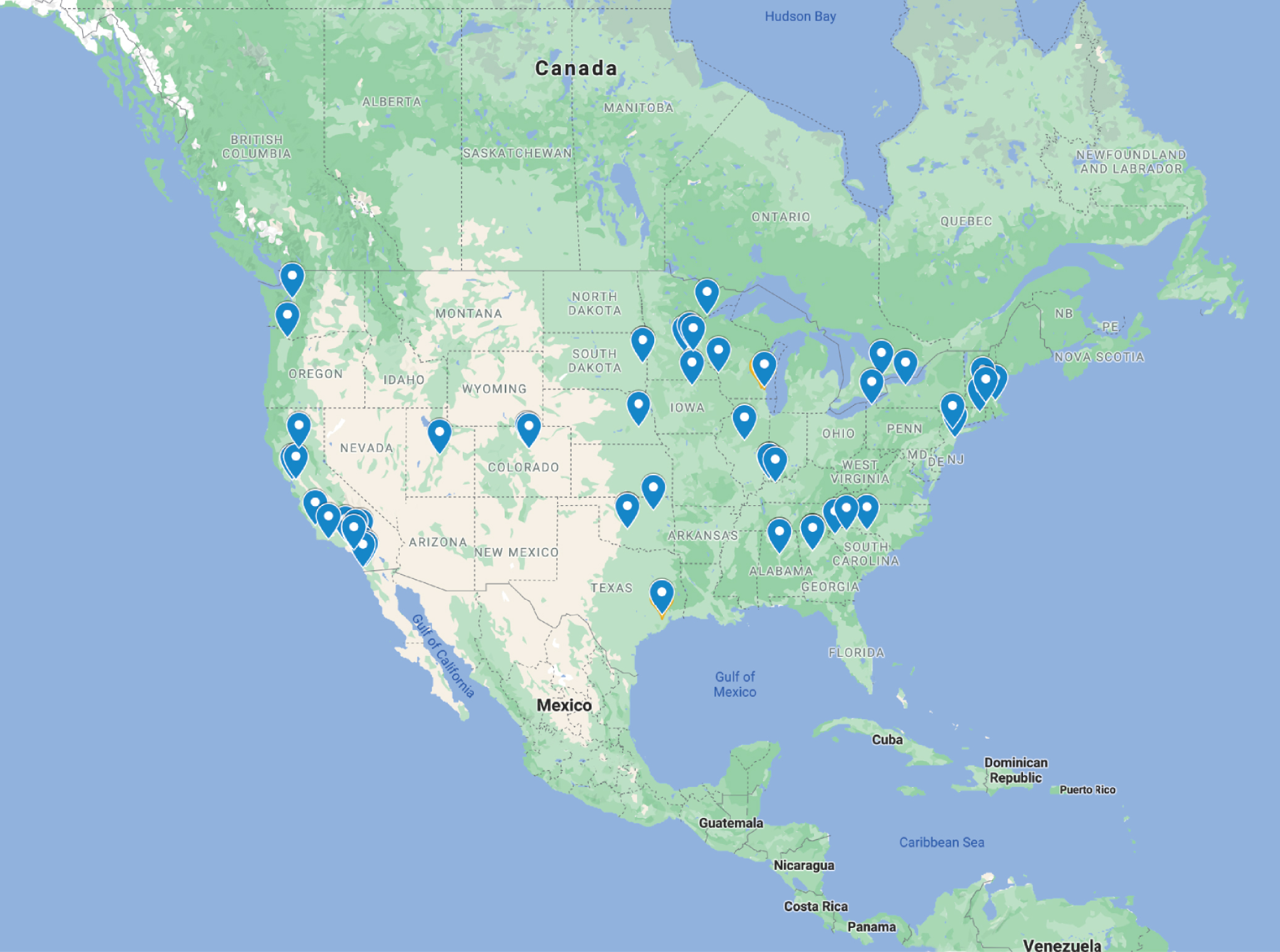
The 2020 Workshop went virtual because of constraints imposed by the COVID-19 pandemic. The main goal of the workshop was to share the knowledge needed to successfully introduce CRISPR technologies at the participants’ home institutions. The virtual format allowed participants from far off places to attend and led to fruitful discussions and collaborations (Figure 1). The workshop allowed educators to come together and devise learning strategies to meet the needs and requirements of learners interested in CRISPR technology. The virtual workshop instilled confidence in educators to utilize available resources in virtual settings. Another insight drawn from the workshop was the possibility to explore virtual resources and online activities to provide context, enhance the hands-on experience, and help reduce reagent/chemical usage. Introduction to these tools highlighted the value of online/virtual resources for biology educators who may have been apprehensive about using them previously.
The use of various virtual learning platforms was well-received by the participants. Instructors and participants used the QUBEShub online platform to communicate, organize material, and submit various documents throughout the workshop (5). Prior to the workshop, participants were given several activities to promote active learning and prepare for the upcoming sessions. These included a walkthrough of a virtual CRISPR lab (via Labster (6)), reading and commenting on an article summarizing the past workshop using Perusall (4, 7), and the creation of an account on the Benchling site that would be used during the workshop. In addition, these activities also provided participants with different mechanisms of interacting with their classrooms in a digital environment. Each participant could also share their concerns and difficulties with the group so that potential solutions to problems could be approached with input from the other participants.
The workshop sessions were 2 hours each day for 5 days, spread over two weeks, via Zoom, which ensured that each session was relaxing for the participants, yet had a strong impact. Participants were also expected to complete assignments between sessions to reinforce the day’s material and prepare them for subsequent activities. Furthermore, sessions were recorded, allowing participants to review material on-demand. The first two live sessions focused on fundamentals of CRISPR biology and the design of a CRISPR-based experiment. Participants learned how to use Benchling to design, evaluate, and select a guide RNA (gRNA), required by the CRISPR-Cas9 system to target a gene for editing. The utility of software applications, like Snapgene, for designing functional expression vector construct(s) for Cas9 and gRNA were also emphasized. Establishing a strong foundation in these areas was important, as many participants had very little formal training in CRISPR related content, if any. The third session gave participants an opportunity to join breakout rooms on various topics, which included the integration of CRISPR in various organisms (plants, invertebrates, vertebrates), the general CRISPR laboratory workflow, improvements required in virtual laboratories for better learning experience (i.e., Labster’s CRISPR module) and the design of Course-Based Undergraduate Research Experiences (CUREs) and/or a Process Oriented Guided Inquiry Learning (POGIL) within a CRISPR framework.
The final two sessions were dedicated to the development of a tangible outcome by the participants. This was organized by breaking the participants up into groups (determined by areas of interest), including introductory-level courses, advanced courses, and online resources, especially virtual labs. The results of these collaborations were presented to all participants on the final day, and all resources were made available to the entire group. This activity enhanced the participants’ motivation towards the design and execution of virtual, as well as hands-on courses on CRISPR.
Despite not being able to engage in a hands-on experience of the CRISPR technology, all the participants concurred that the virtual workshop gave them an experience, which could be relatable to experiences from in-person meetings that they would have attended. Moreover, almost every participant expressed that they would have enjoyed having more sessions. Such feedback was testimony to the fact that learning about CRISPR technology was no longer limited to hands-on laboratory sessions. To enhance the virtual training outcome, incorporation of short laboratory videos with essential technical details of a CRISPR workflow followed by discussions regarding the procedure may perhaps add greater value to future events.
Immediately following the workshop, attendee feedback was solicited via an online survey, regarding the components that attendees were most satisfied with, major learning gains, and recommendations for improvement. We received 54 responses (a 92% response rate) to the survey.
Summary of Immediate Post Workshop Survey Results
It was commonly reported that although attendees were aware of the CRISPR technology, there was a reluctance towards classroom implementation due to a lack of specific knowledge regarding the nuances of the CRISPR workflow and how to deliver it to students. The interactive introductory seminar on the basics of CRISPR garnered positive reviews, where over 90% of first-time attendees indicated they learned new information about both biological and technological aspects of the process (Figure 2).
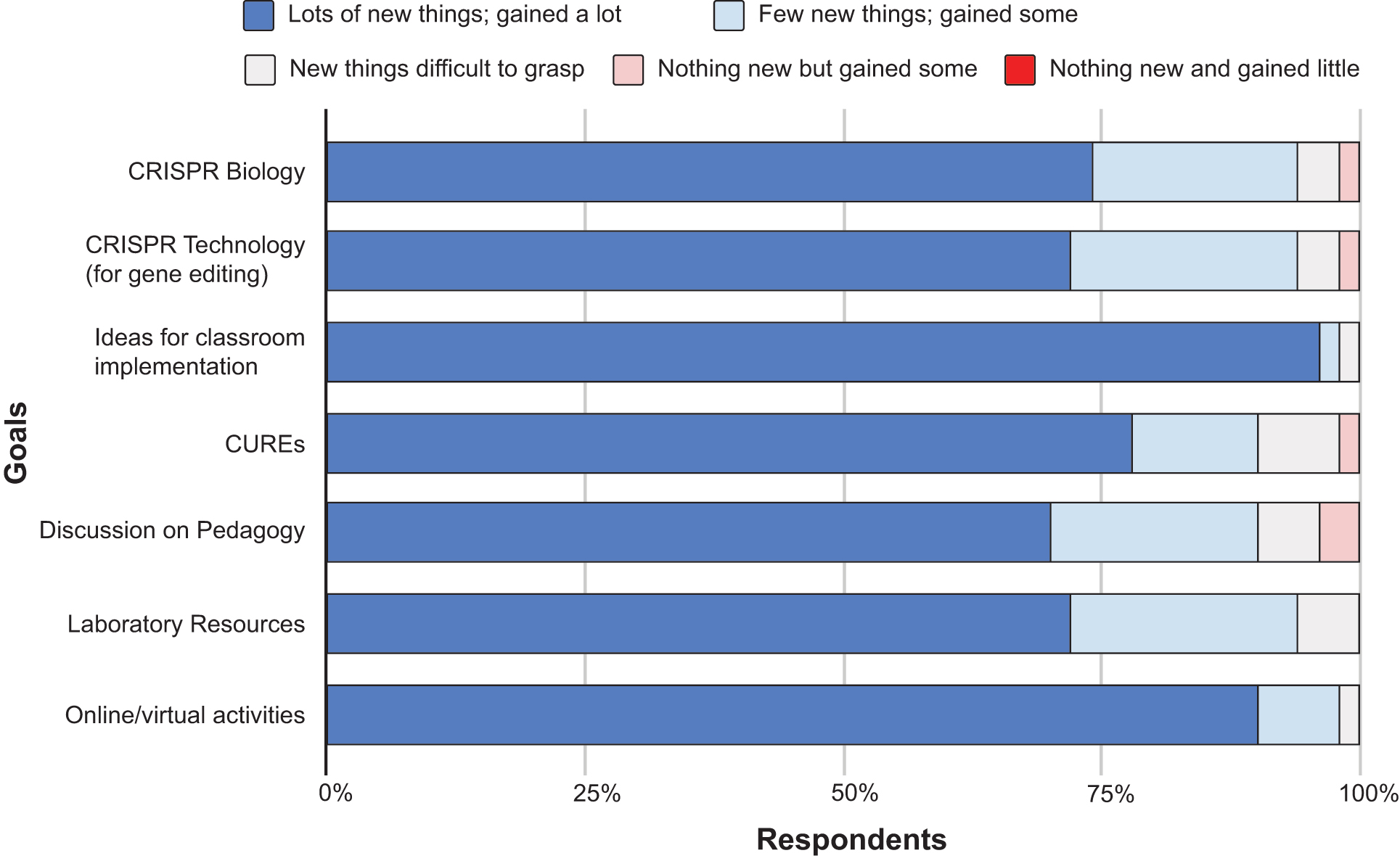
All attendees who responded indicated that the variety of teaching tools used in the workshop (including supplementary videos and virtual activities such as designing guide RNAs) aided in clarifying the concepts. Alongside an increased understanding of the CRISPR process, survey respondents also reported that the workshop provided them with knowledge regarding online resources (e.g., the HHMI Biointeractive tool on CRISPR-Cas9 function and the Benchling tool for gRNA design) that could be used to supplement their teaching. The ability to interact with fellow participants allowed for discussions on learning outcomes, methodologies (e.g., CUREs) already being used to teach CRISPR and potential hurdles for those who are looking to adopt the technology. More than 90% of respondents agreed that these interactions provided them with new ideas for classroom implementation. Additionally, survey results revealed that attendees also found these interactions invaluable in enhancing the level of comfort and willingness to teach CRISPR in addition to providing support networks for any future questions.
Two-day small breakout groups were another major component of the workshop. Groups designed classroom resources (e.g., POGILs and extensive resource lists) to facilitate teaching CRISPR at different student levels (i.e., introductory versus upper year level) and in different model systems. The newly designed resources were shared among all attendees to serve as a starting point for module creation. More than 80% of survey responses found the team activities meaningful for gaining new knowledge. However, there was a consensus among surveyed attendees that a longer time to work on the project would have allowed for greater collaboration and more discussions to occur, leading to a more refined end product. Additionally, it was suggested that each group would have benefitted from having a “CRISPR alumnus” who had already worked with the technology to guide the breakout sessions and help focus the depth of the final product.
The many resources generated were posted in QUBEShub at the end of each workshop day and made available to participants. These resources have been very impactful for attendees and highlight the ability to introduce CRISPR regardless of one’s budget (Table 1). One suggestion was to share new resources created during and after the workshop more widely within the undergraduate teaching community. This suggestion was well received, pending copyright approval from each resource creator.
Table 1. A selection of online resources for introducing and teaching about CRISPR biology and gene editing.
| Online Resource | Source | URL |
|---|---|---|
| CRISPR teaching tools from Genome: Unlocking Life’s Code | Smithsonian Natural History Museum | https://unlockinglifescode.org/crispr-teaching-tools |
| Digital education content, games and activities | Innovative Genomics Institute, Berkeley | https://innovativegenomics.org/education/ |
| Self-paced online course | The Jackson Laboratory | https://learn.education.jax.org/browse/jaxge-online/courses/crispr |
| Biointeractive | HHMI | https://www.biointeractive.org/classroom-resources/crispr-cas-9-mechanism-applications |
Despite increased awareness of the possibilities for teaching CRISPR technology in an online-only format, a concern regarding the lack of hands-on experience was common among attendees looking to implement the technology in a wet lab setting. However, given the challenges with COVID-19, participants acknowledged that this would have to be postponed to a future date. In the meantime, a suggestion was made to have monthly online group meetings for the following one year where attendees could discuss the ways in which they have incorporated CRISPR in their classrooms and seek support regarding any challenges they may be facing.
Overall, 80% of attendees who responded to the post workshop survey indicated that the design and organization of the workshop facilitated their understanding of CRISPR. Since several attendees expressed a desire to incorporate CRISPR into their Fall 2020 courses following this workshop, we polled instructors in an implementation survey about their current and future teaching plans and methodologies used.
Implementation Survey Outcomes
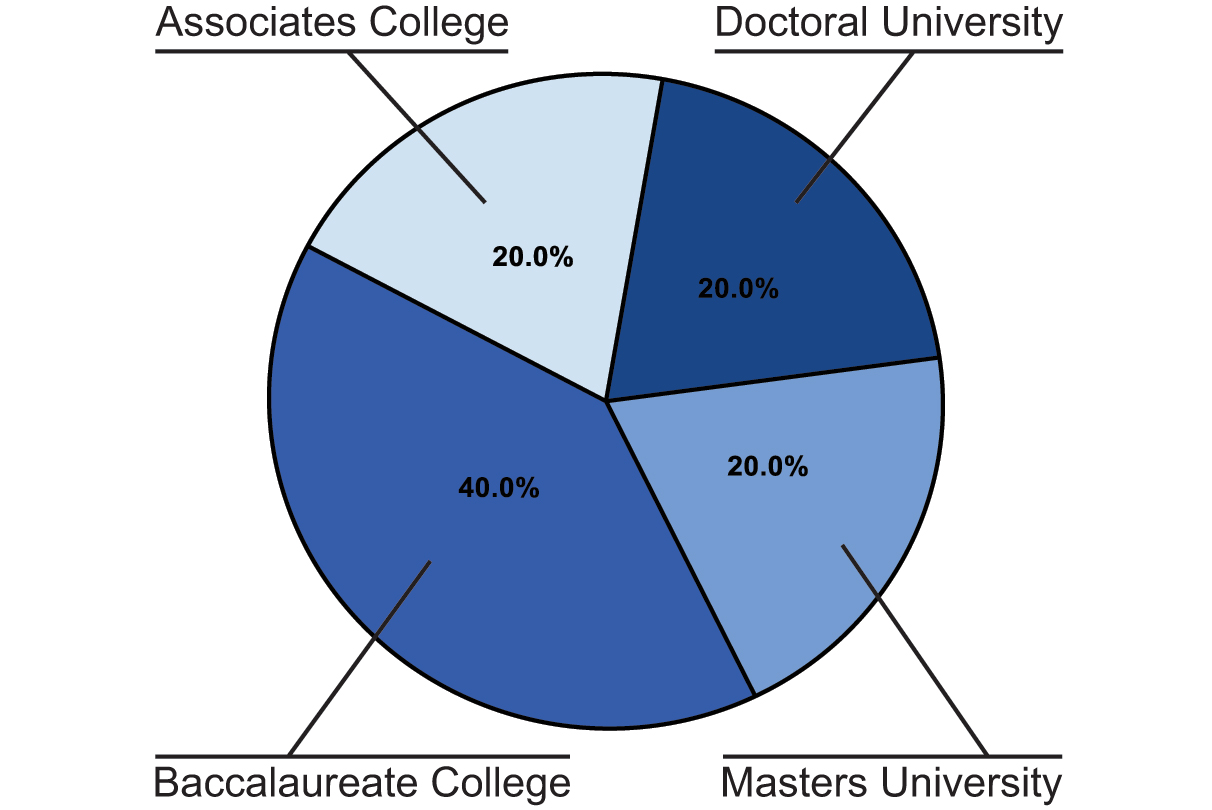
A second survey was sent at the end of the Fall 2020 semester following the summer workshop to understand if and how the workshop helped the participants in their efforts to implement CRISPR in their classrooms. Fifteen workshop attendees (25%) responded to the survey about their plans to implement teaching of CRISPR concepts within their courses. Instructors responded from all different types of institutions (Doctoral Universities, Masters Universities, Baccalaureate Colleges and Associates Colleges), with Baccalaureate Colleges having the greatest representation (Figure 3). Immediately following the workshop, 47% of respondents were already planning to or were highly likely to incorporate CRISPR into their fall 2020 courses. Apart from summer courses, this percentage increased for subsequent semesters (53% for spring 2021; 67% for fall 2021) (Supporting File S1. Virtual CRISPR in the Classroom Workshop–CRISPR lesson incorporation).
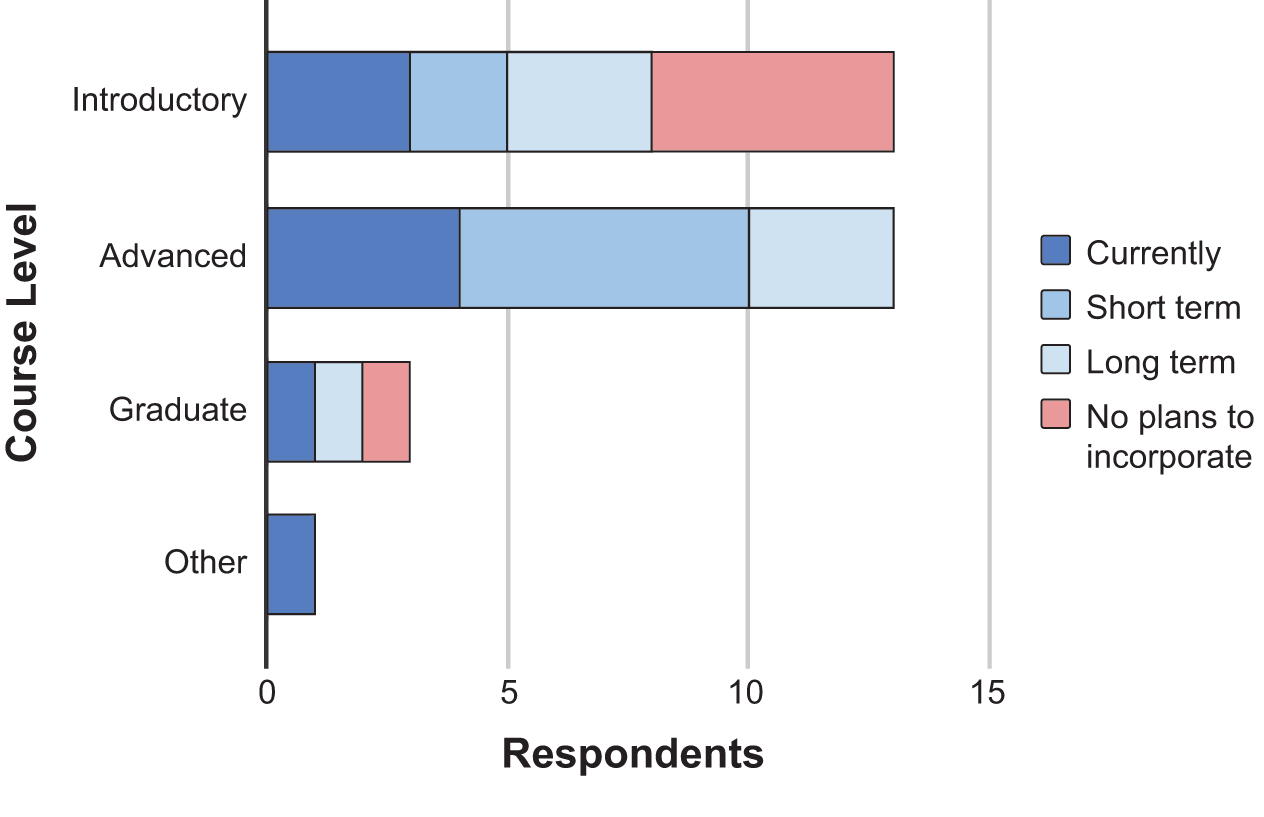
We found that instructors were most likely to add CRISPR into the curriculum in advanced undergraduate courses in the near and long term (Figure 4). All 13 respondents (100%) teaching at the advanced level planned to include CRISPR technologies within the next few semesters. This contrasts with instructors teaching at the introductory undergraduate level, where only 62% planned to include CRISPR technologies in the long term, and 38% of instructors have no plans to incorporate. As nearly half of instructors expressed interest in implementation in introductory level courses, this will be an area of emphasis for future workshops. Only three respondents indicated teaching graduate level courses, and of these, one currently incorporated CRISPR technologies, one had long term plans to include them, and one did not plan to incorporate them into their courses.
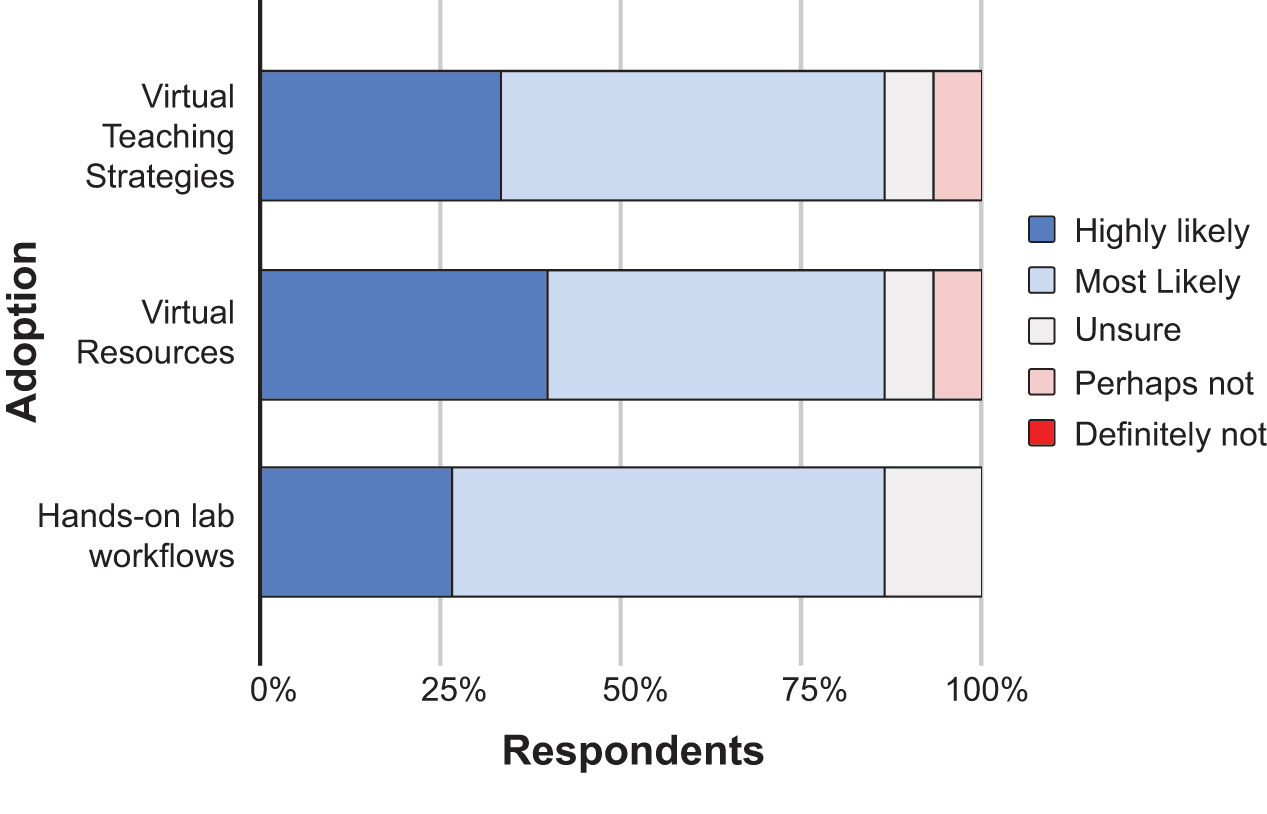
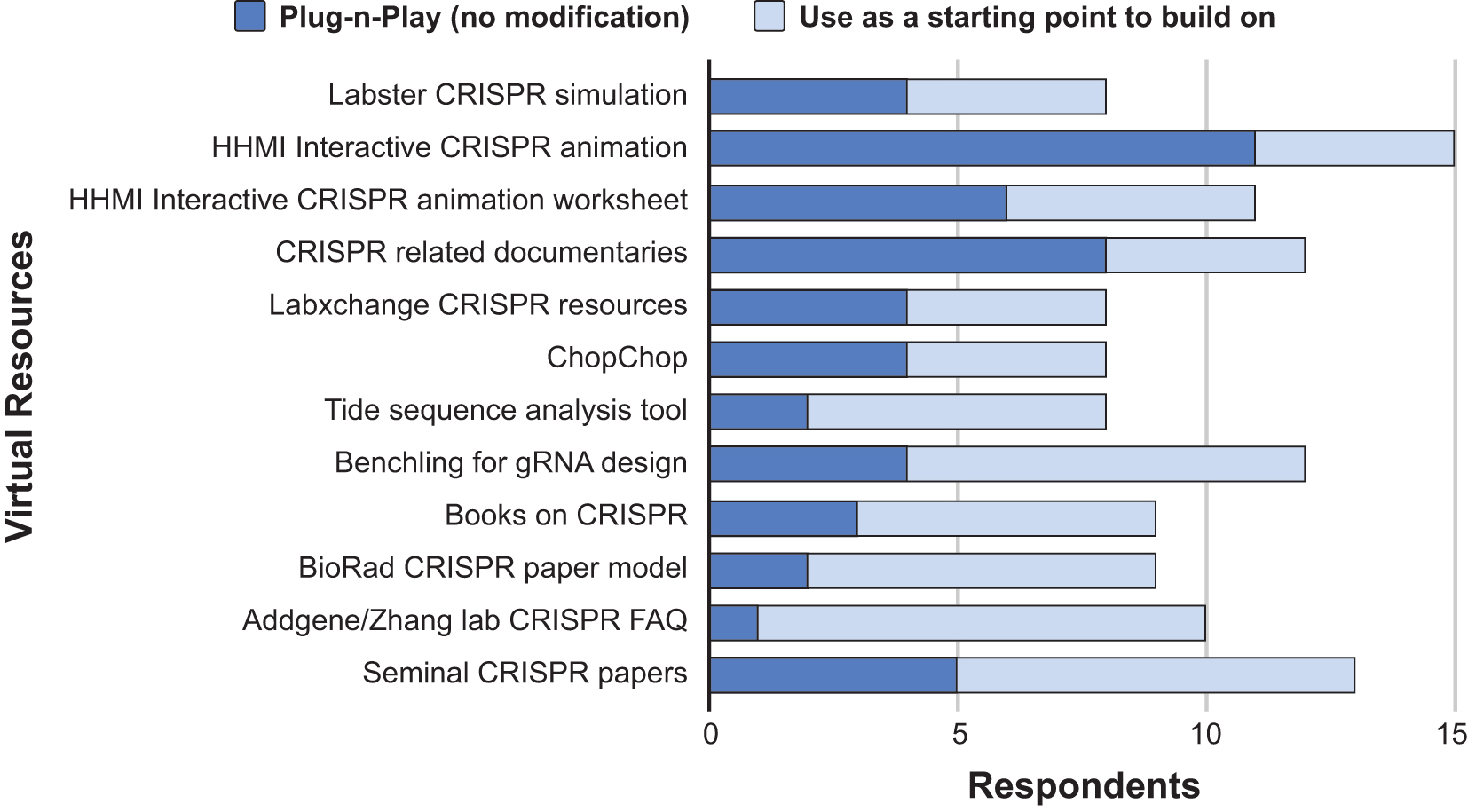
Since the summer 2020 workshop focused mostly on virtual resources to support CRISPR education, we examined the likelihood of participants utilizing virtual teaching strategies, virtual resources, and hands-on lab workflows in their teaching (Figure 5). Given that many instructors taught remotely during the COVID-19 pandemic, it is not surprising that 87% of respondents are “Highly likely” or “Most likely” to employ the virtual teaching strategies or virtual resources highlighted during the workshop. Hands-on lab workflows remain an important teaching strategy with 27% of respondents “Highly likely” and 60% of respondents “Most likely” to incorporate hands-on lab workflows. Looking at adoption of specific resources, instructors indicated which highlighted virtual resources they were likely to use with no further modification (plug-n-play) or as a starting point to build a lesson on (Figure 6). The most widely adopted resource was the CRISPR interactive animation developed by HHMI Biointeractive, which 100% of respondents indicated they planned to use. Other highly popular resources included seminal articles (8, 9), video talks, and using online tools (e.g., Benchling) related to CRISPR technologies.
Incorporation of concepts and technologies related to CRISPR-Cas systems into the classroom laboratory setting will be dependent on the financial resources and time available to instructors at individual institutions. With those constraints in mind, instructors were polled about which hands-on laboratory resources discussed at the workshop they were most likely to utilize. A majority of respondents (80%) indicated that they would utilize an in vitro CRISPR lab workflow (4). Other systems or model organisms represented included roundworm, bacteria, yeast, zebrafish, mammalian cells, or plants. A sizeable number (40%) indicated utilizing one of several commercially available lab kits to introduce CRISPR-Cas systems to their students (Table 2).
Table 2. Commercially available kits for teaching CRISPR technology in the laboratory.
| Kit |
Manufacturer/ Supplier |
EDU Price | URL |
|---|---|---|---|
| E. coli CRISPR Knockout Teaching Kit | abm | $425 | https://www.abmgood.com/i-e-coli-i-crispr-knockout-teaching-kit.html |
| Out of the Blue CRISPR and Genotyping Extension Kits | Bio-Rad | $235-$399 | https://www.bio-rad.com/en-us/product/out-blue-crispr-genotyping-extension-kits?ID=Q0JGD4E08O1Y |
| Exploring the CRISPR-Cas Defense System Kit | Carolina Biologicals | $119 | https://www.carolina.com/dna-models-and-simulations/exploring-the-crispr-cas-defense-system-kit/FAM_211772.pr |
| Using CRISPR To Treat Cystic Fibrosis | Edvotek | $95 | https://www.edvotek.com/135 |
| A-maize-ing Editing: Using CRISPR to Improve Crops | Edvotek | $139 | https://www.edvotek.com/210 |
| Hack the Planet: Using CRISPR to Terraform Mars | Edvotek | $199 | https://www.edvotek.com/310 |
| Knockout! CRISPR Cas Gene Targeting Lab | miniPCR | $215 | https://www.minipcr.com/crispr-cas-knockout-lab/ |
Given the broad and far-reaching nature of CRISPR gene editing technology, we were also interested in how instructors would focus their teaching of CRISPR-Cas systems. The majority (87%) indicated they would spend time on the molecular biology of CRISPR. Emphasis on design of CRISPR experiments (80%), execution of CRISPR-Cas related experiments (73%) and ethics surrounding the use of CRISPR (73%) was also substantial. The fewest respondents indicated they would focus on applications of CRISPR technology (60%).
Focus area: Misconceptions
Since the CRISPR technology is discussed widely in the mainstream and social media, it is not surprising that there are several misconceptions about the technology and its usages. These misconceptions can be used as the basis for designing course content or curricular materials that can be used to achieve student learning outcomes. A few misconceptions that we gathered as part of the workshop and teaching points to address them are given in Table 3.
Table 3. Student misconceptions and teaching points to address misconceptions.
| Misconception | Teaching Point |
|---|---|
| 1. The effects of CRISPR gene editing can be passed on to the next generation. | Students should be reminded of the gene expression level at which CRISPR works and the difference between somatic versus germ cells. Changes made to somatic cells will not affect future offspring. |
| 2. CRISPR technology will lead to the eradication of all diseases that we know of today. | While this might eventually prove to be true, the technology is still in the “early” stages and so researchers are still determining how to use it for disease treatment. It will be a long time before we know which diseases can be treated with CRISPR technology. |
| 3. One of the major advantages of CRISPR is that it can be used by individuals looking to expand their families, to design a child with certain “desirable” characteristics. | This would require a discussion on the ethics relating to CRISPR. It would also be worthwhile to discuss that phenotypic traits can be influenced by a combination of genes, so gene editing for this purpose is not as simple as it seems. The role of environment and its impact on the genotype towards creating a phenotype needs to be emphasized. |
Scientific Teaching Themes
Active Learning
This virtual workshop explored various active learning activities that participants could implement in their courses. Participants now have experienced virtual active learning activities involving CRISPR-Cas systems to help students achieve learning goals through these strategies. Some examples from the workshop include scaffolding CRISPR content to be accessible to non-molecular biologists, doing an interactive walk-through of CRISPR research data analysis, hand drawing a CRISPR experiment workflow (4), and using Perusall software for collaborative article reading and viewing. However, the types of activities and strategies used will vary from instructor to instructor and course to course.
In addition, there is an increasing number of articles on CRISPR/Cas9 technology that are specifically geared towards undergraduate biology educators and classrooms, which can be used as resources. Examples include recent articles by Dahlberg & Groat Carmona (10) and Thurtle-Schmidt & Lo (11).
Assessment
Assessment strategies were discussed throughout the workshop and attending instructors have had a chance to examine and discuss assessment techniques. However, we again expect assessment activities to vary between instructors and courses.
Inclusive Teaching
This report aims to highlight the utility of a virtual workshop to provide professional development and support to a diverse audience of instructors on an emerging topic of importance in biology. Short sessions over multiple days offered free of charge made it possible for an economically, geographically, and institutionally diverse cohort to participate in the proceedings. Workshop content also emphasized cost-effective virtual and hands-on resources for instructors to use to readily incorporate the topic of CRISPR biology into the curriculum at their home institutions at any level of course instruction.
Conclusion
Gene editing using CRISPR-Cas systems is rapidly progressing and transforming both life science research and its applications in medical, agricultural, veterinary, and related fields. As the use of CRISPR technology becomes mainstream, the need to train students widely and to enable their technical skills becomes imperative and can occur by ensuring that educators are conversant with the technology. The effectiveness of these workshops and the need expressed by undergraduate biology educators has motivated the formation of a formal network, which is now supported by an NSF Research Coordination Network grant (Award#2120417 - RCN-UBE: Bringing CRISPR-Cas9 technologies to the undergraduate classroom: an undergraduate instructors' network). While the annual workshops remain the mainstay of the project, additional efforts will be made in the direction of making workshop materials easily available to the larger community of undergraduate biology educators on the QUBES platform and conducting online discussion groups to sustain the momentum generated by the in-person workshops. The stage is now set for sustained growth and dissemination of CRISPR technologies into undergraduate classrooms.
SUPPORTING MATERIALS
Supporting File S1. Virtual CRISPR in the Classroom Workshop–CRISPR lesson incorporation
Acknowledgments
The authors would like to thank QUBEShub for the support provided to host the workshop discussions, and Labster for giving workshop participants access to the CRISPR module. Surveys were conducted with approval from the Hampden-Sydney College Human Research Committee.
References
- Goodsell DS, Autin L, Olson AJ. 2019. Illustrate: Software for biomolecular illustration. Struct Lond Engl 1993 27:1716-1720.e1; doi:10.1016/j.str.2019.08.011.
- Nishimasu H, Ran FA, Hsu PD, Konermann S, Shehata SI, Dohmae N, Ishitani R, Zhang F, Nureki O. 2014. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156:935–949; doi:10.1016/j.cell.2014.02.001.
- Uyhazi KE, Bennett J. 2021. A CRISPR view of the 2020 Nobel Prize in Chemistry. J Clin Invest 131; doi:10.1172/JCI145214.
- Wolyniak MJ, Austin S, Bloodworth LF, Carter D, Harrison SH, Hoage T, Hollis-Brown L, Jefferson F, Krufka A, Safadi-Chamberlin F, Santisteban MS, Soneral P, VanWinkle B, Challa AK. 2019. Integrating CRISPR-Cas9 technology into undergraduate courses: Perspectives from a National Science Foundation (NSF) workshop for undergraduate faculty, June 2018. J Microbiol Biol Educ 20; doi:10.1128/jmbe.v20i1.1702.
- Donovan S, Eaton CD, Gower ST, Jenkins KP, LaMar MD, Poli D, Sheehy R, Wojdak JM. 2015. QUBES: a community focused on supporting teaching and learning in quantitative biology. Lett Biomath 2:46–55.
- Delgado T, Bhark S-J, Donahue J. 2021. Pandemic teaching: Creating and teaching cell biology labs online during COVID-19. Biochem Mol Biol Educ 49:32–37.
- Adams B, Wilson NS. 2020. Building community in asynchronous online higher education courses through collaborative annotation. J Educ Technol Syst 49:250–261; doi:10.1177/0047239520946422.
- Marraffini LA, Sontheimer EJ. 2010. CRISPR interference: RNA-directed adaptive immunity in bacteria and archaea. 3. Nat Rev Genet 11:181–190; doi:10.1038/nrg2749.
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. 2012. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821; doi:10.1126/science.1225829.
- Dahlberg L, Groat Carmona AM. 2018. CRISPR-Cas technology in and out of the classroom. CRISPR J 1:107–114; doi:10.1089/crispr.2018.0007.
- Thurtle-Schmidt DM, Lo T-W. 2018. Molecular biology at the cutting edge: A review on CRISPR/CAS9 gene editing for undergraduates. Biochem Mol Biol Educ 46:195–205; doi:10.1002/bmb.21108.
Article Files
Login to access supporting documents
Kubera-Workshop Report Summer 2020 Virtual CRISPR in the Classroom.pdf(PDF | 815 KB)
S1. Virtual CRISPR in the Classroom Workshop-CRISPR lesson incorporation.docx(DOCX | 182 KB)
- License terms

Comments
Comments
There are no comments on this resource.