Bioinformatics - Investigating Sequence Similarity (Version 4.0)
By Adam Kleinschmit, Benita Brink, Steven Roof, Carlos Christopher Goller, and Sabrina Robertson
Module Description:
The following modules are from a pre-print version of the "Sequence Similarity: An inquiry based and "under the hood" approach for incorporating molecular sequence alignment in introductory undergraduate biology courses" learning resource accepted for publication in CourseSource, which is currently in press.
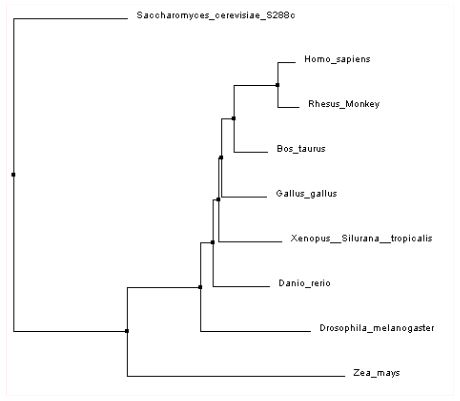
This laboratory module, leads introductory biology students in the exploration of a basic set of bioinformatics concepts and tools. The exercise utilizes simple paper models to help students understand matrices and algorithms prior to use of web-based computational tools. Students start the module by defining sequence similarity and then investigating how similarity can be quantitatively compared between two similar length proteins using a BLOSUM scoring matrix. Students then consider finding local regions of similarity between a sequence query and subjects within a large database using BLAST. Lastly, students practice accessing FASTA formatted sequence information via NCBI databases as they collect sequences for a multiple sequence alignment in order to generate a phylogenetic tree.
Teaching Setting:
This laboratory module was designed for introductory biology students.
QUBES Citation:
Kleinschmit, A.J., Brink, B., Roof, S., Goller, C.C., Robertson, S. (2018). Bioinformatics - Investigating Sequence Similarity. NIBLSE Incubator: Bioinformatics - Investigating Sequence Similarity, (Version 4.0). QUBES Educational Resources. doi:10.25334/Q4VM1P
|