A Short Laboratory Module to Help Infuse Metacognition during an Introductory Course-based Research Experience
Published online:
Abstract
A core competency identified in Vision and Change for undergraduate biology students is the Ability to Apply the Process of Science. Here, we describe a three-week laboratory module for students in an Introductory Cell and Molecular Biology course. The goal of our module is to introduce students to the critical scientific process skill of metacognition early in their undergraduate careers, which is not only important for scientific research, but also for learning new concepts and other types of problem solving. To achieve this, our laboratory module engages students in the investigation of a biological research question while specifically and explicitly prompting students to practice the metacognition regularly employed by scientists. In our research module, students gather information, generate hypotheses, evaluate the utility of different experimental approaches in testing their hypotheses, planning experiments, and analyzing data. In-class and take-home activities prompt students to actively reflect on the information they use to design their experiments and to draw their conclusions. The module has been implemented several times in recent academic years, with two or three concurrent sections of the course taking part each academic quarter. Student evaluations and interviews suggest that this module provides a meaningful introduction to metacognition as it is used in scientific problem solving. Here we present the pedagogical structure of our laboratory module, which could be adapted to engage students in investigating a wide variety of research questions.
Citation
Lee, S.R., Dahlberg, C.L., and Wiggins, B.L.. 2019. A short laboratory module to help infuse metacognition during an introductory course-based research experience. CourseSource. https://doi.org/10.24918/cs.2019.20Society Learning Goals
Science Process Skills
- Process of Science
- Locate, interpret, and evaluate scientific information and primary literature
- Pose testable questions and hypotheses to address gaps in knowledge
- Plan, evaluate, and implement scientific investigations
- Interpret, evaluate, and draw conclusions from data
- Construct explanations and make evidence-based arguments about the natural world
- Address novel questions through authentic research experiences
- Modeling/ Developing and Using Models
- Build and evaluate models of biological systems
Lesson Learning Goals
- Student will participate in science and scientific thinking early in their undergraduate studies.
- Students will design experiments, including choosing appropriate controls.
- Students will engage in metacognition during the process of scientific problem solving.
Lesson Learning Objectives
- Students will be able to evaluate the strengths and weaknesses of data.
- Students will be able to employ prior knowledge in formulating a biological research question or hypothesis.
- Students will be able to distinguish a research question from a testable hypothesis.
- Students will recognize that the following are essential elements in experimental design: identifying gaps in prior knowledge, picking an appropriate approach (ex. experimental tools and controls) for testing a hypothesis, and reproducibility and repeatability.
- Students will be able to identify appropriate experimental tools, approaches and controls to use in testing a hypothesis.
- Students will be able to accurately explain why an experimental approach they have selected is a good choice for testing a particular hypothesis.
- Students will be able to discuss whether experimental outcomes support or fail to support a particular hypothesis, and in the case of the latter, discuss possible reasons why.
Article Context
Course
Article Type
Course Level
Bloom's Cognitive Level
Vision and Change Core Competencies
Vision and Change Core Concepts
Class Type
Class Size
Audience
Lesson Length
Pedagogical Approaches
Principles of How People Learn
Assessment Type
INTRODUCTION
Laboratory courses are often celebrated as hands-on, active learning opportunities. However, many introductory laboratory courses are based on "cookie-cutter" protocols that merely ask students to follow directions, without requiring them to engage thoughtfully with the experiments or observations that they are making. According to the AAAS Vision and Change report (1), the Ability to Apply the Process of Science involves "posing problems, generating hypotheses, designing experiments, observing nature, testing hypotheses, interpreting and evaluating data, and determining how to follow up on the findings" (1). These skills require metacognition, which is the act of critically thinking about one's own thinking (2). Metacognition involves knowledge of how one learns as well as regulation of behaviors that help one learn (3); the latter is particularly crucial for success in scientific research. Moreover, strong metacognitive skills can lead to deeper and longer lasting learning with broader utility in everyday problem-solving (4-6). Because student engagement of metacognitive skills is not often explicitly incorporated into undergraduate courses, despite their importance to learning and the scientific process, we designed a laboratory module that blends a research experience with metacognitive training and implemented it in the last three weeks of an existing ten-week, quarter-long course for introductory biology students.
How is metacognition used in the process of learning and scientific research?
The scientific process involves the critical evaluation of information and tools, continuous monitoring of progress, adjustment of strategies (if necessary), and reflection on results--all of which depend on metacognitive regulation (3,7). Critical thinking about ongoing thought processes allows scientists to assess, evaluate, plan, monitor and apply, and reflect at every stage of the scientific process. These actions are also invoked during active learning (4) and can be prompted in a student research laboratory setting (Table 1) (2,7).

Table 1. Metacognitive processes used by self-regulated learners as applied in scientific research. Processes are adapted from How Learning Works, Ambrose 2010.
Why train introductory undergraduate students in metacognitive practices?
If students are to succeed in advanced laboratory courses, and in research outside of coursework, they need to learn and practice problem-solving skills that are critical to the process of science (1,4). While it often takes years for scientists to internalize and apply metacognitive skills with proficiency, students who are able to reflect on their own activities ("Am I using the correct pipette? Will my action contaminate a sample?") and are able to direct or re-direct their actions ("What is the best next step? How do these data change how I look at this problem?) will have the greatest chance of success in the laboratory (8-10). Yet students who are early in their undergraduate careers are likely to be less able to use metacognitive regulation, even if they have metacognitive knowledge (3). Because metacognition is important for all learning, building these skills may be particularly beneficial for early-career students, regardless of their scientific ambitions. Thus, in order to better prepare students for upper division coursework, the scientific workforce, and greater success in graduate school, metacognitive regulation--including self-reflection and self-direction--should be a part of early scientific training (11,12).
Our module introduces research and metacognitive training to students in their second academic quarter of a Biology curriculum, laying the foundation for continued growth as these students move through college (13). Notably, research experiences--including short, embedded experiences in laboratory courses similar to the short research experience described here--increase student retention in STEM fields and positively affect student psychosocial outcomes (9,14-16). However, many undergraduate students do not have access to research opportunities (12). By including research in an introductory course, we can ensure that our students participate in at least one research project during their college careers.
Overview of the Module
This research module balances the freedom of scientific research with the logistical constraints of an existing course. Here, we provide a framework for meaningful experimental design, experimentation, and analysis at the Introductory level that can run during part of an academic quarter (Supporting File S1). Most significantly, throughout our module, we provide prompts to encourage students to self-reflect ("Where have you seen these genes before? How did you come to your conclusion?") and to self-direct ("What tools will be the best for testing your hypothesis? What would a result of X mean with respect to your hypothesis?") during laboratory section discussions, worksheets, and/or homework exercises. Through the processes described in Table 1, students can generate, revise, and test their own hypotheses. We stress that although the detailed module laid out here describes a specific research topic investigated by students (i.e., control of DNA replication in the model eukaryote, Tetrahymena thermophila), the pedagogical framework of our module could be adapted to accommodate any experimental research question. Tetrahymena thermophila is a student-friendly model eukaryote that has enabled numerous foundational discoveries in molecular cell biology (17,18). As a test of the generalizability of our module, we recently used the same framework to engage students in a place-based research project centered in WWU's backyard, the Bellingham Bay (materials are available upon request). To facilitate adaptation of our module's framework (Supporting File S1) for other topics of interest, we have included both our detailed Tetrahymena-focused materials (Supporting Files S2 and S3), as well as subject-independent versions of our lab manuals and pre-lab lecture presentations (Supporting Files S4-S6 ) that could be adapted to a variety of biological topics.
INTENDED AUDIENCE
This laboratory module was designed for Introductory Cell and Molecular Biology students, at the freshman or sophomore level. At Western Washington University (WWU), this course serves pre-majors in Biology, is a prerequisite for related majors (kinesiology, behavioral neuroscience, biochemistry, etc.), and can be used to fulfill part of the General Undergraduate Requirement. Prerequisites for Introductory Cell and Molecular Biology at WWU include the previous introductory biology course (including some inquiry-based laboratories) and two quarters of general chemistry (limited inquiry-based laboratories). The module is designed to be incorporated into a lecture-and-laboratory course, though it could be reconfigured to a laboratory-only course by introducing more content into the prelab lectures if need be. At WWU, the lecture portion of this course (up to 96 students per lecture section) is taught in concurrent sections by several faculty every academic quarter, while laboratory sessions (24 students each) are led by Master's level graduate student Teaching Assistants (GTAs).
REQUIRED LEARNING TIME
This module takes place over the course of three 3-hour laboratory sessions (Weeks 1-3). Included in these sessions are pre-laboratory lectures and discussions. Additionally, during lecture sessions, students receive a brief (15-30 min) introduction to the relevant molecular pathway under study prior to the need for this information in lab. Students working alone or in small groups are given the option to complete the assigned worksheets and reflections as out-of-class work.
PREREQUISITE STUDENT KNOWLEDGE
To engage most successfully with this module, students should have knowledge relevant to the biological question under study. For example, in the implementation of the module described in detail here, our students examined the impact of overexpression of a DNA replication regulator in a model eukaryotic organism, Tetrahymena thermophila. Through lecture sessions prior to the module, the students were introduced to the general roles that proteins play in cells, how DNA sequence relates to protein sequence, structure, expression levels, and function, and the use of model organisms in research. Students also needed working knowledge of light microscopy and pipetting techniques, which were introduced in laboratory sessions prior to beginning this three-week module.
PREREQUISITE TEACHER KNOWLEDGE
Laboratory instructors need a working knowledge of the tools and concepts required for answering the biological question under study. For example, in our implementation of this module, laboratory instructors needed experience working with the students' light microscopes and familiarity with web-based gene-analysis tools. For the Tetrahymena DNA replication module, our instructors also had a clear understanding of cell cycle progression, regulation, and checkpoints, and familiarity with aseptic technique in handling cell cultures. Detailed lab manuals and preparation protocols, along with a list of needed equipment and materials, have been included here to facilitate instructor implementation of a Tetrahymena module (Supporting Files S2: Laboratory Manual and S3: Tetrahymena protocols); the Tetrahymena educational community can also provide practical knowledge of both Tetrahymena protocols and the implementation of lab modules in undergraduate classrooms (http://faculty.jsd.claremont.edu/ewiley/). To best support student engagement in metacognition, instructors should have training in student-centered teaching practices and the role of metacognition in long-term learning. For introductions to metacognitive training in the biology classroom, as well as its limits, please see the following references: (3,4,19).
SCIENTIFIC TEACHING THEMES
ACTIVE LEARNING
Our 3-week module embedded in a quarter long course connects introductory lecture material with a laboratory experience, and begins training students in self-reflective inquiry that will enhance their metacognition (Supporting File S1). Below, we outline two major facets of our module that rely on active learning: drawing on previous knowledge to tackle new problems and practice with self-reflection and metacognition.
1) Drawing on previous knowledge
In order to reconnect students with content that they have encountered previously, the laboratory "pre-lab" sessions should use familiar images and vocabulary. For our Tetrahymena DNA replication-focused module, the slide(s) that introduce the genes of interest are shared between lecture instructors and the laboratory instructors (Supporting File S7: Tetrahymena Replication Model). The slide is important for implementation of our module because the "alphabet soup" of gene names and gene products is overwhelming and seemingly trivial out of context. Discussion of the basic mechanisms of gene expression and transmission (the "Central Dogma") uses terminology and imagery that students were previously exposed to in the classroom component of this course. While Week 1 (DNA sequence analysis and Bioinformatics) requires students to use new technology to interact with familiar concepts, Week 2 (Introduction to the experimental system) asks students to draw on their existing knowledge of technology (e.g. light microscopes) acquired in earlier laboratory sessions in order to address a new scientific question. Using these scaffolds, students must connect their prior knowledge to their current laboratory experience in order to engage and revise conceptual models.
2) Self-reflection and metacognitive practice
Our Lesson includes activities that require metacognitive practices, but do not directly ask students to reflect on their own metacognition. To prompt students to self-reflect, worksheets provide opportunities for students to address their thinking and changes in mindset post-hoc. During the laboratory sessions, discussions and other activities require cooperation and collaborative thinking by pairs of students. In order to provide students with a platform to describe their ideas and self-reflection on difficulties, laboratory instructors facilitate group discussions surrounding previous data, hypothesis generation, choices in experimental design (controls, experimental assays, etc.), and evaluation of data with respect to the individual and to the group.
To facilitate large group discussions, and in order to prevent any one student being put "on-the-spot", we employ think-pair-share strategies throughout the laboratory modules, especially in Weeks 2 and 3. Students initially brainstorm hypotheses, expected results, and ideal experimental designs. Once student pairs have had time to come to a decision, or have solidified their points of confusion, they are asked to share out, either verbally, or by drawing on the chalkboard. In groups, or as a class, students then reflect on their answers, discuss alternatives, and redirect their thinking or planning to address novel or conflicting points of view.
ASSESSMENT
Laboratory instructors used the following assessments to determine whether students were engaging in self-reflection: laboratory worksheets (addressing content, but also in-laboratory metacognitive practices), in-class discussions and on-the-board reporting of hypotheses and expected results, and questions on homework that ask for individual reflections on decision making and previous knowledge. Questions on the final laboratory practical exam assessed whether students had learned how to use instrumentation correctly. To assess content knowledge, some classroom instructors used exam questions on their final exams. These questions assessed whether students were able to describe (very briefly) their data and conclusions that might be drawn from it. Findings on the success of the module in terms of student impacts are discussed briefly in the Teaching Discussion and at length in a separate manuscript (26).
INCLUSIVE TEACHING
The demographics of students enrolled in WWU's Introductory Cell and Molecular Biology course from Spring 2015-Winter 2018 included: 30% first-generation college students (defined as having two parents who did not complete college), 71% low-income FAFSA applicants, and 16.4% students who identify as belonging to a traditionally underserved racial minority group (Hispanic/Latinx, African American/Black, Native American/Alaskan Native, Native Hawaiian/Other Pacific Islander, or Filipino/Hmong/Vietnamese). It is known that many students, especially those from the groups listed above, leave science not because they lack the potential to succeed, but because they do not identify with those in the scientific community or view themselves as capable of success as scientists (12,20). By infusing explicit practice with metacognition in the context of a research module, we aim to have all of our students become aware of their engagement as legitimate scientists with diverse experiences and points of view.
To increase the sense that a scientist identity is achievable by our students, we explicitly connect metacognitive processes that are critical for science (listed in Table 1) to the laboratory and metacognitive module activities that the students engage in. Moreover, the opportunity to engage in a true scientific struggle allows students to reframe difficult moments in science as a natural part of doing real science rather than a referendum on their own potential as scientists, potentially increasing student development of a growth mindset towards science. In our implementation of this module as a Tetrahymena-focused research project, we were also able to introduce relevant preliminary data that had been obtained by another undergraduate student researcher (21). In sum, students can make the connection that research can be difficult, that previous research can be improved upon, and that they can be the ones to improve upon it.
The structure of this module requires the equal participation of all students, with evidence of their involvement collected in the form of various assessments, including worksheets and participation in class discussions. Students are given time to process concepts and organize their own thoughts through take-home work or in small-group explorations before being asked to report out to the class or to complete hands-on laboratory activities. This provides students with the opportunity to fully participate in and bolster their confidence for in-class work. The metacognitive prompts and assigned homework are designed to be inclusive of diverse points of view, allowing for a variety of possible answers (see Supporting Files S8-S10). Finally, we emphasize the importance and strength of a diverse community to making scientific progress in our module. Student pairs must draw on each other's strengths to accomplish several distinct tasks, from collecting information in silico to execution of laboratory protocols to reflecting on their experiences and decision-making.
LESSON PLAN
Our laboratory module integrates into a lecture and laboratory combination course (Introduction to Cell and Molecular Biology) and includes pre-lab lectures, laboratory practice, and discussions (Table 2). The biological question investigated by students in our implementation of this module is how the cell cycle is regulated. We used the single-cell ciliate organism Tetrahymena thermophila because it is a relatively inexpensive eukaryotic cell model system and is conducive to simple microscopy and genetic manipulation (17). Through a collaboration with the Ciliate Genome Consortium, we obtained a strain of Tetrahymena that can overexpress the protein Cdt1 fused to Yellow Fluorescent Protein (YFP). Because CDT1 is required for licensing origins of replication (preparing them for the initiation of DNA replication during S-phase) (22), it likely plays a central role in cell-cycle control in Tetrahymena; preliminary data obtained by a student in a course taught by a CGC member supported this possibility (www.suprdb.org/index.php/searchDetails/geneName/cdt1). As noted above in the Overview of our module, we have also included more general materials for instructor use (Supporting files S4-S6) that could be adapted to address research questions that differ from the Tetrahymena-focused one we used in the implementation we describe here in detail.
Over the module's three lab sessions, students combined their existing scientific and technical knowledge with Tetrahymena-specific experimental assays to develop and address an unanswered scientific question using the CDT1 overexpression strain (Table 2). Our laboratory sections met one time per week, as described below. Sessions could be run more often, though time should be allotted to ensure that students can finish the accompanying worksheets. All laboratory sessions began with a 10-15 minute pre-laboratory presentation (given as a PowerPoint slideshow) and discussion (prompted by phrases, questions, or data in the PowerPoint) led by the laboratory section GTA. Students worked as pairs in the laboratory and followed their laboratory work with a post-laboratory activity or homework assignment completed outside of class. Our students participated in the module at the end of the academic term, in part so that they could apply skills in using a light microscope and micropipetting acquired in prior weeks. However, the module could be implemented wherever it fits best with existing curriculum (provided students are given at least an introduction to the use of micropipettors and light microscopes), so we describe the timeline in terms of first, second, and third weeks of the module.
WEEK 1 (LAB)
The pre-laboratory 1 presentation includes a review of the Central Dogma, the genetic code, and potential impacts of gene mutation on a gene product (Table 2, Supporting Files S7 and S11). Students are also introduced to gene analysis bioinformatics tools accessible online (Clustal W, EXPASY, Blast and Pfam) in the presentation (Supporting File S11). During the laboratory session students explore the structure and functions of provided "unknown" human gene sequences (CDT1 and its partner proteins, Supporting File S7) using bioinformatics tools (Supporting File S8); student work here is guided by explicit direction in their lab manual (Supporting File S12) as well as by hands-on support from their lab instructors. Loss-of-function mutations in any one of the genes that encode these proteins result in the human genetic disorder Meier-Gorlin syndrome (22-24), so sequence files containing these gene variants are also included for student analysis (Supporting File S13). To directly engage student metacognition in preparation for the next two laboratory sessions, students complete a worksheet which contains questions that require students to assess their existing knowledge about the genes they have examined and to evaluate how their previous knowledge informs their ability to make an experimental prediction (Supporting File S8, Table 1 - Assessing and Evaluating). For example, students are asked to make a prediction for how overexpression of a Cdt1 homolog might impact a Tetrahymena cell and then answer the question "What makes you think that?" Students are also prompted to reflect on the utility of specific bioinformatics tools that they used in lab and the value of research performed in model organisms rather than humans. To further lay the foundation for hypothesis generation and experimental planning in Weeks 2 and 3, students use the Tetrahymena genome database (25) to identify and explore the potential functions of Tetrahymena homologs of the human genes they examined. As an optional assignment, students can be asked to formally present their sequence analysis data in the form of a scientific figure (Supporting File S14); as part of this assignment, students are again asked to reflect on the value and motivation for their analysis, while also making an experimental prediction. With up to 12 pairs of students working in one laboratory section, at least two groups can be assigned to (or choose) each gene, so that students can compare their findings, and, in essence, replicate each other's findings.
WEEK 2 (LAB)
The pre-laboratory presentation focuses on Tetrahymena as a research model organism and a review of DNA replication and the cell cycle (Table 2, Supporting File S15). The pre-laboratory discussion actively engages students in assessing, evaluating and the early stages of planning (Table 1 - Assessing, Evaluating, Planning) by having them brainstorm what types of experiments could be done in Tetrahymena to study the function of genes of interest. Specifically, in a manner guided by the instructor, students share their initial predictions for the biological impact of overexpression of one of those genes, CDT1 in Tetrahymena, given what they know about DNA replication control and Cdt1 protein function (Supporting File S7). In principle, instructors could choose any genes in a pathway for which there are Tetrahymana strains that have been modified to overproduce the encoded protein; we recommend exploring a single gene and collecting data in aggregate as a class. As a class, students also assess and evaluate the findings and strengths and weaknesses of preliminary data obtained on CDT1 overexpressing Tetrahymena cells. These data are publicly available at on a database of student-generated data set up by the Ciliate Genomics Consortium (www.suprdb.org/index.php/searchDetails/geneName/cdt1). Discussion points can include: importance of reproducibility, sample size and complementary methods of experimental analysis in being able to draw conclusions about biological pathways from scientific studies.
Finally, in preparation for Week 3, the Week 2 laboratory activities encourage mastery of gross phenotypic and behavioral observations, cell counting, nuclear staining, cell imaging, and taking measurements of cell and nuclear size using live and fixed wild-type Tetrahymena (Supporting File S2). These activities lay the foundation for students to engage in scientific planning (Table 1 - Planning), as it builds their knowledge of the experimental tools that they will have at their disposal for Week 3 to test the hypotheses that they have identified in Week 2. To engage students in scientific planning further, students complete a worksheet following the laboratory session that requires reflection on the nature and quality of data they obtained from the experimental assays they had learned in Week 2 (Supporting File S9). As students begin to connect an initial hypothesis for how CDT1 overexpression might impact Tetrahymena cells to assays that they could use to test their hypothesis, they must assess the specific strengths and weaknesses of different experimental approaches. Additional metacognitive prompts are included to encourage students to explicitly assess what knowledge or information they used to refine their hypotheses, their level of confidence in their decision making, and what types of information might boost their confidence (Table 1 - Applying and Monitoring, Reflecting).
WEEK 3 (LAB)
The pre-laboratory presentation centers on what is involved in building a testable hypothesis, including the initial research question, a prediction for outcomes of an experimental test, and what variables must be considered (Table 2, Supporting File S16). Though engagement with the pre-laboratory discussion, students refine their research questions, establish hypotheses for testable outcomes of CDT1 overexpression, and plan the protocols to test their hypotheses, based on the experimental assays they learned and reflected on in Week 2 (Table 1 - Assessing, Evaluating, Planning). After implementing their plans in lab, students enter their raw data into spreadsheets, which are compiled by the GTAs, and outside of class, students prepare data reports (Supporting File S10). As part of their data reports, students include a discussion of whether the data supported or failed to support their hypotheses and why they have made the conclusions that they have (Table 1 - Applying and Monitoring, Reflecting). Students are also prompted in the data report assignment to reflect on the impact data aggregation made to their confidence in their conclusions compared to the individual measurements students made with their laboratory partners and the preliminary data originally obtained by others. This prompt is aimed at solidifying concepts of reproducibility and sample size in the scientific process.
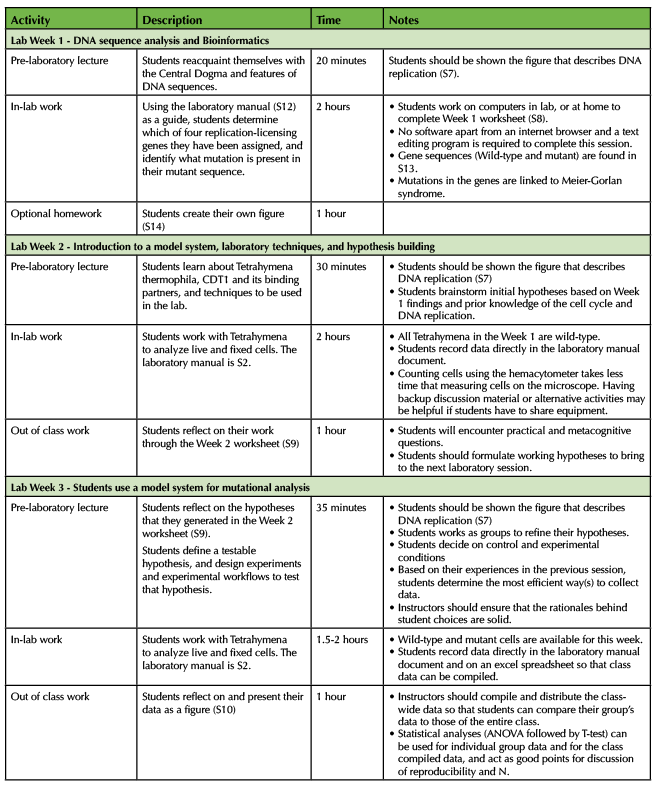
Table 2. Short Research Module - Teaching Timeline
TEACHING DISCUSSION
SUMMARY HIGHLIGHTS FOR OUR STRUCTURE
Our research module introduces the process of science to students through a collaborative research project that depends on activities designed to directly evoke and build the following metacognitive skills: accessing prior knowledge, hypotheses generation, refinement and revision, and reflection on outcomes. These activities implicitly direct students to assess, evaluate, plan, monitor, reflect, and share throughout the three-week scaffolded research project. In this way, students engage fully in the metacognition that scientists actively use in the course of research.
Because our module targets introductory students, we hope that providing this framework encourages students to further strengthen their own identities as scientists and members of the scientific community. Students are encouraged to think about how their data fits in (or doesn't) with previous results, and how their data compares to that of their immediate peers. In our implementation, by drawing on student-generated preliminary data from another university, students also experience the scientific process as a community effort and an evolving process. In this way, they experience first-hand how data and data interpretation can change current models of cell biology in particular, and of scientific theory in general.
CONCLUSIONS FROM ASSESSMENTS OF OUR MODULE
Our module has three overarching learning goals: 1) to introduce the process of science and scientific thinking to students who are early in their science careers, 2) to guide students to explore how experimental design fits into the process of science, and 3) to lead students to value the use of metacognition in the process of scientific research. One type of impact assessment performed was to ask for feedback about the laboratory module itself through open-ended "exit questions" at the end of the module (26). Student responses often included statements that show value placed on metacognition in the research experience (Table 3). For example, students reported "figuring out" methods, "think[ing] about what we were doing," and getting familiar with instruments and "how I could use them." While no student directly stated that they were using metacognition, many of them referred to the metacognitive processes shown in Table 1.
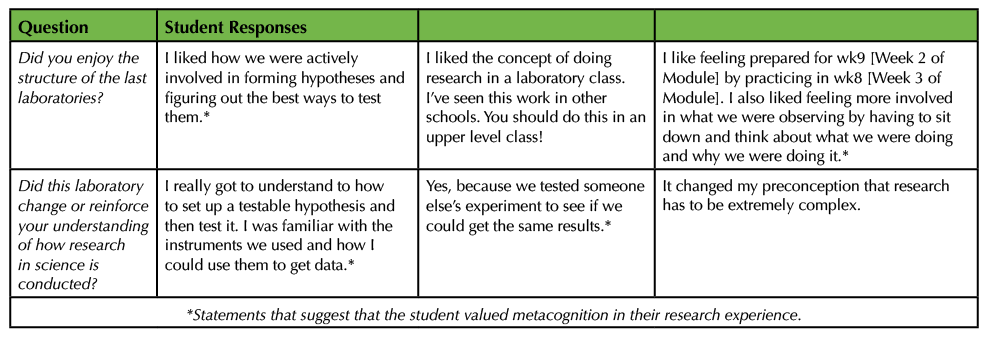
Table 3. Student responses to questions about the authentic research module
Furthermore, we interviewed students after their Introductory Cell and Molecular Biology experiences as part of a formal focus group study. Students repeatedly reported that laboratory experiences are more engaging when they are asked to make choices in their experimental design, even if these are very simple choices (such as choosing which two of three conditions to compare). Providing students with choice gives them a sense of agency in the laboratory, and if justifications for a choice are required, may cause students to reflect more thoughtfully on their decisions. These types of responses suggest that our learning goals may have indeed been achieved through this module.
CONSIDERATIONS FOR THE FUTURE
Our experiences and preliminary data point towards the importance of opportunities for choice and revision to most effectively engage students in the laboratory (26). Some laboratory sections brainstormed possible follow-up experiments which ranged from technical troubleshooting to testing new hypotheses and why such experiments might be warranted. In our focus group discussions, students reported that being allowed to make bad choices but then to revise their decisions in a subsequent laboratory session would provide strong learning opportunities. While our module didn't provide students with the opportunity for revision due to time constraints, future modules that allow students to implement their follow-up experiments would allow students to engage more deeply with the iterative metacognitive processes that are integral to research. If time allowed, it could be beneficial to design an entire fourth and fifth week of the module dedicated to troubleshooting, building a class-wide data set, and data analysis.
Because the in-class discussion component of our module is important, laboratory instructors should be well-versed in student-centered teaching and learning techniques to best support student-driven decision-making and metacognition. In our implementation, we relied on graduate teaching assistants to lead the laboratory module. Based on our experience, we highly recommend that training for teaching assistants include an introduction to metacognition and the role it plays in the scientific process and long-term learning. It may also be beneficial for laboratory and classroom instructors to explicitly discuss the parallels between principles of self-directed learning and the process of science.
Our module aims to be inclusive of diverse students by engaging students in explicit metacognitive practice so that they can identify as scientists, soliciting individual student perspectives through diverse activities and assignments, and emphasizing the power of collective data in science. We envision expanding this module to include exposure to the work of scientists of diverse backgrounds, engaging students in a reflection on what this means for who can become a scientist.
ADAPTATIONS FOR OTHER INSTITUTIONS AND PROJECTS
Our three-week module provides a framework for explicitly engaging Introductory Biology students in the metacognitive practices regularly employed in science. Because the framework for our module is built around introducing students to the process of science, this framework is highly adaptable; we describe a few possible adaptations and ideas for finding new research questions below.
As mentioned in the Overview above, we recently tested the adaptability of this module to other research topics by piloting a place-based research project, looking at the effects of low-light culturing on algae samples from Bellingham Bay. To facilitate the adoption of the framework of this module by other instructors with specific research interests, please see the subject-independent "generic" templates of our materials (Supporting files S4-S6). Instructors who are interested in adapting our module for research organisms other than Tetrahymena can use the Bioinformatics worksheet (Supporting file S6).
Tetrahymena thermophila are relatively simple organisms to grow, maintain, and visualize with simple light microscropy, making them an ideal model organism for introducing students to research. The protocols that are included in this article are generally simple, of low cost and non-toxic (Supporting Files S2 and S3). For instructors hoping to find other transgenic Tetrahymena genes that can be overexpressed or knocked out to use in a similar experiment, we suggest browsing the SUPRdb website (http://suprdb.org/index.php/experiment/showall) as a starting place (27). SUPRdb is a clearinghouse for unpublished results from the Tetrahymena community, many of which were generated by undergraduates. Therefore, an added bonus to building on findings reported on SUPRdb is that students can be introduced to the fact that undergraduates are capable of generating new knowledge and that science is, at its heart, a collaborative endeavor. We also have benefitted from a partnership with the Ciliate Genomics Consortium for finding strains and discussing protocols (http://faculty.jsd.claremont.edu/ewiley/). Similar partnerships with other professional societies or research groups can also be fruitful sources of un- or under-characterized research organisms and strains.
SUPPORTING MATERIALS
- S1. Short Research Module - Overview figure of different implementations
- S2. Short Research Module - Laboratory Manual
- S3. Short Research Module - Tetrahymena protocols
- S4. Short Research Module - Generic Prelabs (bundled)
- S5. Short Research Module - Generic Laboratory Manual
- S6. Short Research Module - Generic worksheets
- S7. Short Research Module - DNA Replication Slide
- S8. Short Research Module - Week 1 worksheet
- S9. Short Research Module - Week 2 worksheet
- S10. Short Research Module - Week 3 figure assignment
- S11. Short Research Module - Week 1 Prelab lecture
- S12. Short Research Module - Week 1 Sequence analysis manual
- S13. Short Research Module - Gene Sequences and mutations
- S14. Short Research Module - Sequence analysis figure assignment
- S15. Short Research Module - Week 2 Prelab lecture
- S16. Short Research Module - Week 3 Prelab lecture
ACKNOWLEDGMENTS
We thank the Ciliate Genomics Consortium, and specifically Emily Wiley and Doug Chalker, for Tetrahymena strains and their support of undergraduate teaching assistants in the preparation of our module (funding from NSF IUSE grant DUE1431837 to EW). We are also grateful for the support and input of colleagues and collaborators: Hannah Jordt, Leah Lily, David Leaf, Jose Serrano-Moreno, Anna Groat-Carmona, and Lynn Pillitteri. This work could not have been accomplished without the invaluable efforts of our graduate and undergraduate student teaching assistants: Tian-Qing Yen, Mariah Colton, Trevor Bloom and Bailey Jochim.
References
- AAAS. 2011. Vision and Change in Undergraduate Biology Education >> Final Report.
- John D. Bransford Editor. 20000801. How People Learn: Brain, Mind, Experience and School (Expanded Edition). Washington, DC, USA National Academies Press.
- Stanton JD, Neider XN, Gallegos IJ, Clark NC, Tomanek D. 2015. Differences in Metacognitive Regulation in Introductory Biology Students: When Prompts Are Not Enough. CBE--Life Sci Educ 14:ar15.
- Tanner KD. 2012. Promoting Student Metacognition. CBE-Life Sci Educ 11:113-120.
- Cook E, Kennedy E, McGuire SY. 2013. Effect of Teaching Metacognitive Learning Strategies on Performance in General Chemistry Courses. J Chem Educ 90:961-967.
- Peter C. Brown. 2014. Make it stick: the science of successful learning. The Belknap Press of Harvard University Press, Cambridge, Massachusetts.
- Brownell SE, Freeman S, Wenderoth MP, Crowe AJ. 2014. BioCore Guide: A Tool for Interpreting the Core Concepts of Vision and Change for Biology Majors. CBE-Life Sci Educ 13:200-211.
- Shortlidge EE, Bangera G, Brownell SE. 2017. Each to Their Own CURE: Faculty Who Teach Course-Based Undergraduate Research Experiences Report Why You Too Should Teach a CURE. J Microbiol Biol Educ 18.
- Cooper MM, Kerns TS. 2006. Changing the Laboratory: Effects of a Laboratory Course on Students' Attitudes and Perceptions. J Chem Educ 83:1356.
- Cooper KM, Soneral PAG, Brownell SE. 2017. Define Your Goals Before You Design a CURE: A Call to Use Backward Design in Planning Course-Based Undergraduate Research Experiences. J Microbiol Biol Educ 18.
- President's Council of Advisors on Science and Technology (PCAST). 2012. PCAST Documents & Reports | The White House. Engage Excel Prod One Million Addit Coll Grad Degrees Sci Technol Eng Math.
- Bangera G, Brownell SE. 2014. Course-Based Undergraduate Research Experiences Can Make Scientific Research More Inclusive. CBE-Life Sci Educ 13:602-606.
- Harrison M, Dunbar D, Ratmansky L, Boyd K, Lopatto D. 2011. Classroom-Based Science Research at the Introductory Level: Changes in Career Choices and Attitude. CBE-Life Sci Educ 10:279-286.
- Gregerman SR, Lerner JS, Hippel W von, Jonides J, Nagda BA. 1998. Undergraduate Student-Faculty Research Partnerships Affect Student Retention. Rev High Educ 22:55-72.
- Russell SH, Hancock MP, McCullough J. 2007. Benefits of Undergraduate Research Experiences. Science 316:548-549.
- Henter DIH Justin Nicholes, Fang-Yu Liao, Aaron Beasley, and Heather. Short-Term Research Experience (SRE) in the Traditional Lab: Qualitative and Quantitative Data on Outcomes | CBE--Life Sciences Education.
- Ruehle MD, Orias E, Pearson CG. 2016. Tetrahymena as a Unicellular Model Eukaryote: Genetic and Genomic Tools. Genetics 203:649-665.
- Collins K. 2012. Perspectives on the ciliated protozoan Tetrahymena thermophila. Methods Cell Biol 109:1-7.
- Dye KM, Stanton JD, Tomanek D. 2017. Metacognition in Upper-Division Biology Students: Awareness Does Not Always Lead to Control. CBE--Life Sci Educ 16:ar31.
- Hurtado S, Cabrera NL, Lin MH, Arellano L, Espinosa LL. 2008. Diversifying Science: Underrepresented Student Experiences in Structured Research Programs. Res High Educ 50:189-214.
- SUPRdb | Welcome.
- Bicknell LS, Bongers EMHF, Leitch A, Brown S, Schoots J, Harley ME, Aftimos S, Al-Aama JY, Bober M, Brown PAJ, van Bokhoven H, Dean J, Edrees AY, Feingold M, Fryer A, Hoefsloot LH, Kau N, Knoers NVAM, Mackenzie J, Opitz JM, Sarda P, Ross A, Temple IK, Toutain A, Wise CA, Wright M, Jackson AP. 2011. Mutations in the pre-replication complex cause Meier-Gorlin syndrome. Nat Genet 43:356-359.
- Bicknell LS, Walker S, Klingseisen A, Stiff T, Leitch A, Kerzendorfer C, Martin C-A, Yeyati P, Al Sanna N, Bober M, Johnson D, Wise C, Jackson AP, O'Driscoll M, Jeggo PA. 2011. Mutations in ORC1, encoding the largest subunit of the origin recognition complex, cause microcephalic primordial dwarfism resembling Meier-Gorlin syndrome. Nat Genet 43:350-355.
- Burrage LC, Charng W-L, Eldomery MK, Willer JR, Davis EE, Lugtenberg D, Zhu W, Leduc MS, Akdemir ZC, Azamian M, Zapata G, Hernandez PP, Schoots J, de Munnik SA, Roepman R, Pearring JN, Jhangiani S, Katsanis N, Vissers LELM, Brunner HG, Beaudet AL, Rosenfeld JA, Muzny DM, Gibbs RA, Eng CM, Xia F, Lalani SR, Lupski JR, Bongers EMHF, Yang Y. 2015. De Novo GMNN Mutations Cause Autosomal-Dominant Primordial Dwarfism Associated with Meier-Gorlin Syndrome. Am J Hum Genet 97:904-913.
- TGD | Tetrahymena Genome Database Wiki.
- Dahlberg CL, Wiggins BL, Lee SR, Leaf D, Lily LS, Jordt H, Johnson T. 2019. A Short, Authentic Course-based Research Module provides Metacognitive Benefits in the Form of More Sophisticated Problem Solving. J Coll Sci Teach 48(4):22-30.
- Wiley EA, Stover NA. 2014. Immediate Dissemination of Student Discoveries to a Model Organism Database Enhances Classroom-Based Research Experiences. CBE-Life Sci Educ 13:131-138.
Article Files
Login to access supporting documents
A Short Laboratory Module to Help Infuse Metacognition during an Introductory Course-based Research Experience(PDF | 189 KB)
S1. Short Research Module-Overview Figure.pptx(PPTX | 38 KB)
S2. Short Research Module-Laboratory Manual.doc(DOC | 7 MB)
S3. Short Research Module - Tetrahymena Protocols.docx(DOCX | 397 KB)
S4. Short Research Module - Generic Prelabs.pptx(PPTX | 350 KB)
S5. Short Research Module - Generic Laboratory Manual.doc(DOC | 7 MB)
S6. Short Research Module - Generic Worksheets.docx(DOCX | 22 KB)
S7. Short Research Module - DNA Replication Slide.pptx(PPTX | 40 KB)
S8. Short Research Module - Tetrahymena Week 1 Worksheet.docx(DOCX | 15 KB)
S9. Short Research Module - Tetrahymena Week 2 Worksheet.docx(DOCX | 503 KB)
S10. Short Research Module - Tetrahymena Week 3 Figure Assignment.docx(DOCX | 15 KB)
S11. Short Research Module - Tetrahymena Week 1 Prelab Lecture.pptx(PPTX | 75 KB)
S12. Short Research Module - Week 1 Sequence Analysis Manual.pdf(PDF | 2 MB)
S13. Short Research Module - Gene Sequences and Mutations.docx(DOCX | 160 KB)
S14. Short Research Module - Sequence Analysis Figure Assignment.docx(DOCX | 15 KB)
S15. Short Research Module - Week 2 Prelab Lecture.pptx(PPTX | 3 MB)
S16. Short Research Module - Week 3 Prelab Lecture.pptx(PPTX | 989 KB)
- License terms

Comments
Comments
There are no comments on this resource.