The following sessions will be offered on Monday, July 24. Sessions will be 2 hour long, hands on introductions to resources and pedagogical approaches. There will be three rounds of sessions (10:30am, 2:00pm, 6:30pm) and many of the presenters will be available throughout the week. An overview of the sessions in a sortable, searchable table can be found here.
Avida-ED: An artificial life platform for teaching evolutionary principles and how to "do" science
Description:
Would you like your students to engage with evolutionary mechanisms using a digital model that is not just a simulation of the effects of evolution? Are you interested in implementing an active learning and inquiry-based scientific experience in your course?
Using the free browser-based digital evolution platform Avida-ED your students can design experiments and collect data to observe how random mutations and evolution in action can produce complexity. This authentic research experience addresses common misconceptions in an engaging and exploratory manner that many of our students find to be “really cool and helpful,” and “a vivid way to learn” about evolution. For many of our students this experience was the first time they identified with the scientific community: “[With] the Avida-ED project, [it] felt like I was a researcher.”
In our session participants will perform a set of exercises designed to address common misconceptions regarding the process of evolution. We’ll then describe the independent research projects carried out by groups of students using Avida-ED. Our session’s initial exploration of this curriculum will provide the basis for a discussion about how workshop participants might implement Avida-ED lessons and/or research projects in their own courses.
This Avida-ED curriculum has been implemented in biology courses at Michigan State University and around the US, ranging from high school through advanced undergraduate. We will also present data on work we have done to assess the effectiveness of Avida-ED implementations on student learning of evolutionary concepts and science process skills.
For more information about Avida-ED click here.
Resources for this session can be found in the Collections area [linked here]
Integrative Cases for Evolution Education
Room C104Date & Time:
Monday, July 24th
6:30 pm
Description:
Many biology instructors find evolution a difficult topic to teach and many students find evolution a difficult topic to learn. Part of this difficulty stems from the complexity of the evolutionary process that requires knowledge spanning across biological subdisciplines to fully understand. The Evo-Ed (http://www.evo-ed.org) project provides resources to help educators teach evolution using integrative examples of trait evolution. Pre- and post-course assessments indicated that students who learned evolution in this context were better able to explain the molecular basis of mutation, describe how mutations lead to phenotypic change and make mechanistic links between genotypes and phenotypes. In this interactive BioQUEST workshop session, participants will learn about the integrative cases of trait evolution. By the end of the session, participants will be able to implement one or more integrative case in a classroom setting. Stemming from this session, participants will work on developing resources that can supplement and build upon this approach to evolution education.
Resources for this session can be found in the Collections area [linked here]
For more information about Evo-Ed click here.
The ESTEEM collection: Computational tools to support modeling and analysis of biological systems at the introductory level and beyond.
Description:
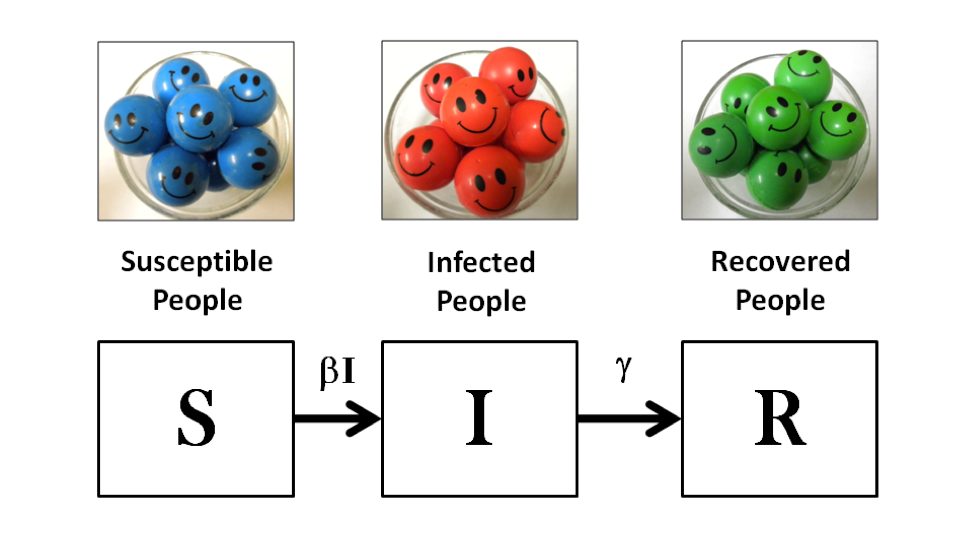
In this session, I will introduce the Biological ESTEEM Collection, an online suite of Excel-based modules that facilitate mathematical exploration of a wide range of biological concepts. These modules are designed to guide early learners step-by-step through the reasoning that underlies a particular model or analysis, while advanced users can modify the default calculations to better reflect the actual features of a given biological system. Today, we will focus on two modules: (1) Island Biogeography, in which students determine which biotic and/or abiotic factors best predict species diversity in discrete habitat patches; and (2) SIR BuildIt, in which students first model the spread of a generic infectious disease, then refine the model to reflect more precisely the properties of a specific disease of their choice.
Resources for this session can be found in the Collections area [linked here]
Data Science in the Undergraduate Biology Classroom using R
Description:
Data science is becoming an increasingly important curricular topic to include in our undergraduate biology classrooms. A biology major, however, already has to gain some proficiency in chemistry, physics, calculus, and statistics - so where are we going to fit another proficiency such as data science, and how different is data science from statistics anyways?
In this session I will focus on the benefits and challenges of teaching the statistical software R in undergraduate biology or biostatistics courses. I will discuss in particular some of the ways QUBES software tools can be used to bring computation and analysis using R into the classroom. I will introduce (1) Jupyter Notebooks, (2) RStudio, and (3) Serenity, a beta R Shiny web application being designed specifically for classroom use. These programs include tools that follow the concept of literate programming introduced by Donald Knuth, which describes documents that combine exposition, data analysis, and visualizations, leading to a reproducible human-readable report of the data analysis performed.
We will then go deeper into the many tools and services made available in the R universe for students to learn and practice data science, including online courses, R-Markdown reports, data dashboards, and interactive Shiny apps. Finally, we will explore some example data sets (or even your own!) using Serenity, which combines the usability of common point-and-click statistics packages, like JMP and SPSS, with the power and programming of R.
After attending this session, participants will be invited to create data science modules using one of these tools. This session is open to faculty with all levels of experience with R.
Data Nuggets: Bringing authentic research and data in the classroom to unearth students’ quantitative and inquiry skills
Description:
Data Nuggets are targeted classroom activities focused on developing quantitative skills for K-16 students. They bring cutting edge science and data into the classroom, helping students develop a deeper understanding of quantitative reasoning in the context of science. In addition, scientists can use Data Nuggets to share their research with broad audiences. Each Data Nugget includes a dataset from real contemporary research for students to graph, interpret, and use when constructing an explanation.
In this session, participants will learn strategies to best utilize this valuable classroom resource and have the opportunity to develop a Data Nugget of their own. With a focus on climate change data, we will provide access to an online source of freely available published data that can be used in classroom inquiry projects. We will guide participants through development of a Data Nugget with this dataset, modelling a classroom experience where students ask their own questions, use data to support their claim, and create a report to communicate their findings. Data Nuggets produced during this session will be hosted on our website (http://datanuggets.org/search-current-data-nuggets/) and on QUBES Hub (https://qubeshub.org/groups/datanuggets). Alternatively, participants are invited to bring data from their own research. If you choose to write a Data Nugget from your own research, please come prepared with the data selected, analyzed, and a research question in mind. We will be available throughout the Summer Institute to assist participants in revising and refining their Data Nuggets.
Beagle Investigations Return with Darwinian Data
Description:
Too often the Galapagos Finches are presented as a canonical example of evolution without providing students with the opportunity to engage with data or the types of reasoning that biologists use to make sense of their similarities and differences. This problem space provides a collection of introductory materials and data resources designed to support students as they reason about the evolutionary relationships between the species.
Participants will learn about BIRDD and problem spaces, become oriented to the Galapagos as an ecological and evolutionary laboratory, and explore multidimensional collections of Finch morphology data using Serentiy (a beta R Shiny web application being designed specifically for classroom use). Participants will then write a report using best practices for data management and reproducible research, informally share their research, and then delve further into a project of their choosing.
BIOMAAP: Biology Students Math Anxiety and Attitudes Program
Description:
The current quantitative reasoning skills of our students, and their willingness to engage withnew quantitative content, can be key roadblocks in teaching modeling in the undergraduate biology classroom. Students’ mathematics anxiety and lack of confidence can lead to decreased performance, in a cycle where students with negative math experiences are then reluctant to engage with new quantitative content, leading to further negative experiences. Fortunately, explicit strategies to increase student comfort with quantitative content can help students successfully engage with the use of mathematical models in biology. Much of the recent biology education reform has emphasized conveying the value of quantitative skills to our students (i.e. the central role computation, statistics, and modeling play in modern scientific discovery), in the hopes that this will increase student motivation. However, student motivation is not dependent solely on valuing the task. The other key component is expectancy: how likely is one to succeed at this task?
In this workshop, participants will experience a variety of short, easily adoptable materials to address mathematics attitudes in biology students by increasing students’ expectations of success. These materials were developed for the NSF-funded Biology Students Math Anxiety and Attitudes Program (BIOMAAP). The general approaches include fostering a growth mindset, using metacognition (thinking about their own process of doing math) to refine student efforts, avoiding stereotype threat, and developing foundational numeracy skills. We will also provide a brief background on the different approaches and current evidence of their effectiveness. The materials are targeted at introductory undergraduate biology courses, but are not biology-content-specific and are thus appropriate for a range of introductory and upper division undergraduate courses. Materials include both out-of-class and in-class activities that can be led by an instructor or peer mentor, and actively engage students in reducing their own mathematics anxiety and improving their attitudes toward mathematics.
An Invitation to Modeling: Exploring the process of science through the process of modeling
Room C102 Description: Do you teach models such as Hardy Weinberg Equilibrium or population growth? Do your students understand how these models are used to explore biological concepts? We, as an interdisciplinary group of biologists, mathematicians, and education researchers, have been using these models, ubiquitous in biology curricula, as an example of how to introduce biology students to the process of modeling. In keeping with our emphasis on better understanding through multiple representations, we offer a multi-faceted approach to understanding common biology models, beginning with a hands-on activity and emphasizing how to use graphical representations of numerical data to build a verbal representation and the corresponding symbolic representation. Students who lack the background to understand the formulas can nevertheless learn to think mathematically through the interplay among experience, data, and graphs, and activities can support the understanding of models, while students with various mathematical exposure and adapted to courses with varying levels of mathematical expectation. We introduce the framework using outlines of potential HWE and population growth activities. Therefore, there is room to contribute to the more complete development of these activities. There is also the ability to apply this framework to enrich the teaching of other models in biology. Choose your own adventure: Each participant can choose among the following three activities to explore our modeling framework. HWE Lactase Persistence Activity HWE PopGen Activity Resource-Limited Population Growth Activity
Description: The double helical model of the structure of DNA (Deoxyribonucleic acid), published in 1953, revolutionized biology. Watson and Crick developed this model by integrating key experimental findings from chemistry, physics and biology. The model immediately suggested mechanisms for genetic information storage and use of the genetic blueprint of life, leading to the central dogma of biology. Today, using interdisciplinary approaches we continue to explore and understand the subtleties of DNA structure and function that enable storage of information, exquisite regulation of individual genes in response to various signals from within and outside the cells, and much much more. Understanding DNA structure and function has also inspired scientists to engineer and design new molecules that do not exist in nature. This workshop will begin with a hands-on demonstration of how Watson and Crick developed the double helical model of DNA and take the story forward to the first few X-ray structures of DNA. In the remainder of the workshop participants will interactively explore the structures of DNA and its various complexes using the RCSB Protein Data Bank website (www.rcsb.org), tools, and resources to learn about DNA structure and function. In addition to visualizing structures online, participants will also make paper models of DNA that they can use for teaching and discussions about DNA. Educational materials, and the foundations for various case studies will be made available. We hope to encourage faculty to develop case studies and/or exercises that teachers and students around the world can use to learn about nucleic acids and proteins in 3D. For more information about DNA and related Educational resources go to pdb-101.rcsb.org and search for DNA in the top search-box.
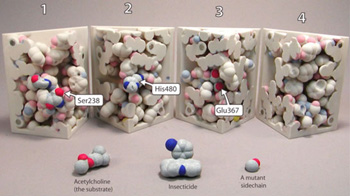
Description: HHMI BioInteractive offers a wealth of resources for teaching how to analyze data. Our particular strength is in combining engaging media that tell compelling science stories with ready to use data analysis activities using real data. If you need to teach fundamental statistical analysis, you will find many tools at our main website including Excel spreadsheet tutorials, a guide to basic statistics, and an interactive tutorial for developing an intuitive understanding of sampling, normal distribution, and standard error of the mean. Our advanced resources are also available from HHMI BioInteractive QUBES page, that was developed by the Faculty Mentoring Network from last year. Some of our resources can be found on our partner sites at Science in the Classroom and The National Center for Case Study Teaching in Science. At the end of our session, we invite you to discuss ideas for how to make our resources fit introductory biology courses better, and suggestions on what content we can develop to fill a need. Description: While we all agree that math and quantitative thinking is critical for biology majors, biology curricula at most institutions remain resolutely math-free. The reason is often the lifelong fear or distaste for mathematics of many biology majors, which is often intentionally reinforced by biology faculty. Much of this fear and distaste arises from the abstract nature of mathematics, and the challenge of building the intuition of how mathematics applies to biology. Based on the long history of BioQuest and my own experience, I have found that the best steps for helping biologists develop an understanding of the role of math in biology include: (1) physical manipulatives, (2) computer simulations, (3) derivation of mathematical relationships from core principles, and (4) analysis of real data sets (Jungck et al. 2010). This workshop will focus solely on the first element, physical manipulatives. Manipulatives help learners break through prior fears and then develop an appreciation for how mathematical reasoning informs problem solving, inference, and precise communication in biology. Adding manipulatives to quantitative teaching is the biological equivalent of a laboratory portion of the course. Two main concerns that are often voiced as challenges to implementing the use of manipulatives is that the tasks are often game-like, and some students will consider the tasks to be beneath them. Additionally, the use of physical props seems antiquated in comparison to modern computer gaming. However, it is interesting that neither of these concerns is applied to the labs that are frequently used to teach introductory biology! There the tried and true methods for teaching the scientific method, the core concepts of biology, etc., include the use of some of the exact same experiments today that emeriti professors did in their undergraduate days! This workshop will focus on teaching the use of manipulatives by working through the exercises. Additionally, there will be discussion about the appropriate courses and sections that each could be applied. Finally, we will conclude with a discussion of the mathematics needed for each exercise and how these topics can be provided to the math faculty for use in the required mathematic classes. Reference: Description: This workshop will use a variety of models (available for loan from the MSOE Model Lending Library) to explore how random variations within a gene result in a phenotype of insecticide resistance in mosquitoes. We will use a case study approach that will enforce a conceptual understanding of a variety of topics in the context of preventing the spread of Zika virus: We will explore these topics as we discuss the use of insecticides that target acetylcholinesterase, the enzyme that recycles the neurotransmitter acetylcholine. We will also utilize molecular landscapes created by David S. Goodsell to place the molecular machines we discuss in the context of their location within the body. Throughout the workshop we will address the value of hands-on, minds-on physical models in uncovering student conceptions and taking them to the next level of understanding, as well as effective ways of incorporating models in the classroom. For more information about the Center for BioMolecular Modeling, go to: http://cbm.msoe.edu/ Resources for this session can be found in the Collections area [linked here]
Description: Homeostasis and the role of hormones in maintaining homeostasis are fundamental concepts covered in Introductory Biology courses. In this workshop, we will use the example of the hormone erythropoietin (EPO) which is essential for regulating red blood cell (RBC) production to introduce participants/students to modeling. Participants will be introduced to a mathematical model and will be provided the corresponding VPython script that they would alter to simulate different scenarios. We will follow-up with a reflection on the guiding principles for the design of the activity, the challenges of implementing the activity in a multi section/multi instructor introductory course and finally invite participants to discuss ways in which they may incorporate similar activities in their classroom. Description: Many scenarios in the life sciences can be modeled with discrete difference equation models which simulate how quantities change over evenly spaced intervals. The construction and analysis of models of discrete difference equations do not require knowledge of calculus, and thus make excellent exemplary models in courses where the typical student may not be proficient in calculus. This session will introduce a few discrete difference equations with a wide variety of applications in modeling population growth/decline and harvesting/stocking, patient drug dosing, and population genetics. Session participants will be provided with materials developed for an undergraduate freshman level course on discrete mathematical modeling for the life sciences including readings, lecture slides, and computer lab projects. Though the materials have been developed for a math modeling course, they would be appropriate for adaptation to a variety of biology courses. Workshop participants wishing to incorporate discrete difference equations models in their courses will be encouraged to apply for participation in the Discrete Modeling for Life Science QUBES Faculty Mentoring Network (aka the Discrete FMN) where participating biology faculty will engage in converting the materials presented in this workshop into single modules, separated from the developmental sequence of a math course, to be utilized in the participating faculty’s Fall biology courses. During the Fall semester, the Discrete FMN will provide support for the implementation and assessment of the newly developed modules. Resources for this session can be found in the Collections area [linked here] Description: How many kinds of models can we put in a case. This workshop will introduce finding and adapting cases using visual models, mathematical models, 3D models, kinesthetic models, visual models and simulations. Each participant will identify a case, adapt the learning objectives to their own class and find different kinds of models to insert into case activities.
Description: To get the most out of this workshop, participants should come prepared to develop a set of classroom activities that ask students to model their ideas. Participants should select a biological concept/model that they would like to work with. We will be working with the definition of a scientific model as - “a set of ideas that describe a natural process (Cartier 2001).” So, in choosing a concept/model, participants should keep this mind. In class, we will be using examples based on primary literature articles dealing with co-evolution, food chains, competition, predator prey dynamics. Other concepts could include almost any other biological phenomena such as natural selection, phylogenetic trees, genetics, photosynthesis, cell division etc. Description: We developed an alternative to the usual “Calculus for Life Sciences” course that is a gateway course for biology freshmen and sophomores. There is widespread dissatisfaction with the standard calculus course (no modeling, no biology, too technical, single variable only, linear equations only). All the leading voices in biology education (NSF, NAS, HHMI, AAMC) have called for a very different class, teaching dynamics using differential equations, in which the differential equations are developed as models of biological and physiological systems, and are studied by numerical integration using Euler’s method, as well as qualitative analysis (such as equilibrium points, oscillations etc.) Our course presupposes only high-school algebra, and develops the necessary math by putting the concept of modeling first, and then showing how a model, described in terms of stocks and flows in positive and negative feedback relations, gives rise to a “change equation”, which assigns “change vectors” to each point in the system’s state space. The evolution of the system is then given by following the change vectors everywhere from a given initial condition. In our session we will briefly outline this approach, do some modeling exercises, and answer lots of questions, including many of the form “how the heck can you do this with freshmen without calculus?” We have been giving this course at UCLA for the past 4 years; this past Fall we registered 1100 students. We will present data on the reception of the course, which has been very positive. We’ve written a textbook, called “Modeling Life: the Mathematics of Biological Systems” that will be published by Springer in August.
Description: Case It! is a project providing a framework for collaborative case-based learning via free, open-ended molecular biology simulations. This project had its genesis at the 1995 BioQUEST workshop and is currently part of the Science Case Network. Existing cases are based primarily on genetic and infectious disease, but the software has wide application in the classroom and is also useful for course-based research projects. Case It v6 is a computer simulation that performs standard laboratory procedures and produces realistic results using any DNA or protein sequence. Capabilities of the software include electrophoresis, multiplex PCR, Southern, Western, and dot blotting, ELISA, and SNP or expression microarrays. Case It v6 is integrated with stand-alone and web-based bioinformatics tools for multiple sequence alignment and phylogenetic tree-building. The software can also directly access online databases for the analysis of DNA or protein sequences associated with cases. Participants in this session will use Case It software to create ‘virtual gels’ as models for actual laboratory results. They will also use Case It in conjunction with the EVO-ED ‘monkey opsin’ case, to show how this case can be expanded to enable undergraduates to conduct original research on color vision in primates, utilizing bioinformatics techniques. There will be an opportunity to explore other features of Case It that may be useful for developing additional modeling activities for the classroom. Resources for this session can be found in the Collections area [linked here] For more information about Case It! click here.
Description: Stochastic processes are those in which a certain level of unavoidable randomness prevents the outcome from being predicted with 100% certainty. Such stochasticity is not just intrinsic (to varying degrees) within all physical processes, but in many cases directly motivates specific practices in scientific research; e.g., the use of experimental replicates and controls, formal hypothesis testing, and replication and meta-analysis of others’ results. Unfortunately, life science undergraduates across different levels of academic achievement and class standing often struggle to understand this core concept. As a result, they are unlikely to see the value in many standard research practices: after all, if we expect a 3-to-1 phenotypic ratio for a given cross, why spend the time performing dozens of matings when just four offspring seem sufficient? Developing a deeper appreciation for the interplay between deterministic and stochastic processes — between the signal and the noise in a given data set — is seldom the main focus of a given study. As a result, students may take years to gradually extract this hidden subtext from their lab exercises and research projects. However, strategic simulations (Jungck and Calley 1985) offer a computational approach in which students directly confront the role of stochasticity as they collect and analyze simulated data at a greatly accelerated pace, essentially compressing the research cycle to better align with the schedule of a typical undergraduate laboratory. In this session, drawing on the framework of strategic simulations, we will: Resources for this session can be found in the Collections area [linked here] 
Date & Time
Monday, July 24th
2:00 pm and 6:30 pm
In this session, we will lead participants through a HWE hands-on activity based on lactase persistence SNP data from Morales et al. (2011). We developed this activity to make the process of modeling explicit, help the students connect and reflect upon what they learn in math and biology courses, and give the model context that enriches learning.
This activity expands on a typical discussion of evolutionary forces to explore using HWE as a null model. The hands-on entry point is the use of the PopGen simulator (http://www.radford.edu/~rsheehy/Gen_flash/popgen/) to experiment with how various evolutionary forces can affect allele frequencies over time.
Most students have seen an exponential model for population growth and know that these models predict unlimited growth. Better models, such as the logistic growth model, should incorporate assumptions to account for limitation of resources. In this activity, participants collect data from a physical simulation of resource-limited growth, prepare a variety of graphs to visualize the data, construct a verbal model based on their observations and the graphs, and (for those who want to go this far) construct a symbolic model from the verbal model. Resources for this activity are located here.
To the Double Helix and Beyond: Exploring DNA Structure and Function in 3D
HHMI BioInteractive
“Beanbag” biology: feelings count!
Jungck, J. R., Gaff, H., & Weisstein, A. (2010). Mathematical manipulative models: In defense of “beanbag biology” CBE Life Sciences Education, 9, 201–211.
Evolution in Action: Using Models to Explore the Molecular Basis of Insecticide Resistance
Design and implementation of a modeling activity in an Introductory Biology course
Let’s do it discretely! An introduction to discrete difference equations models in the life sciences
Mutating Cases with Models
Using web-based technology to promote student use and understanding of mathematical and visual models
Teaching Differential Equation Modeling to Biology Freshmen
Case It: Integrating molecular biology computer simulations and bioinformatics into case-based learning and student research projects
Introduction to Stochastic Modeling