Software
- Details
- Published on Wednesday, 07 January 2015 18:30
Resources: Software
Find software
Featured Software
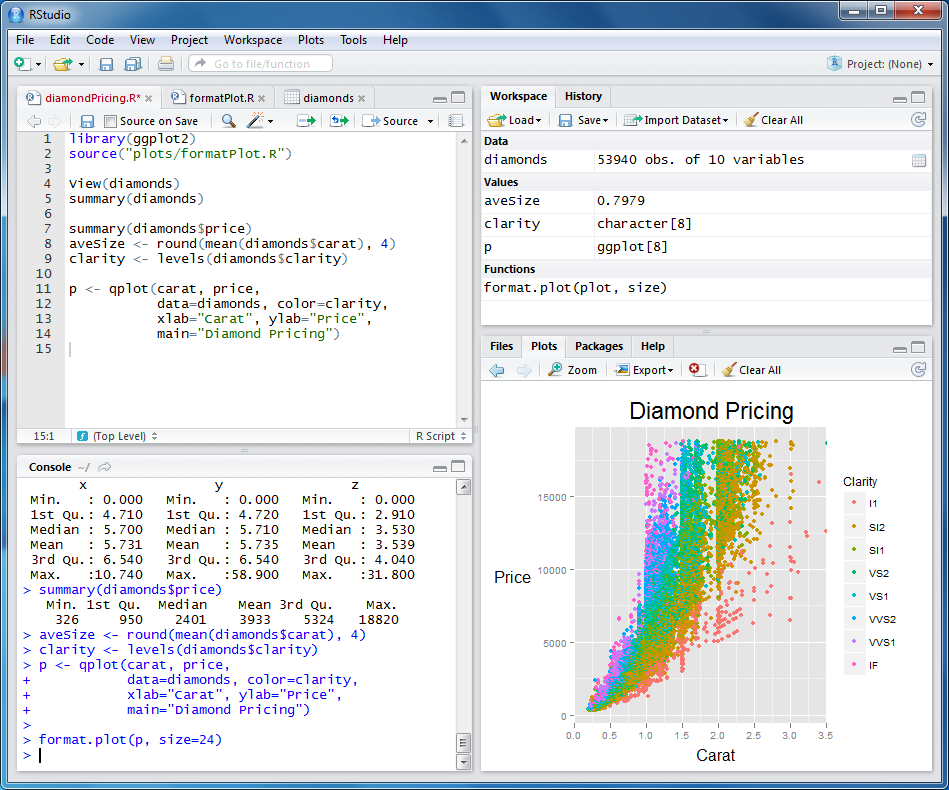
R-Studio IDE for R
RStudio is a GUI for R, the statistical programming language.
Links of interest:
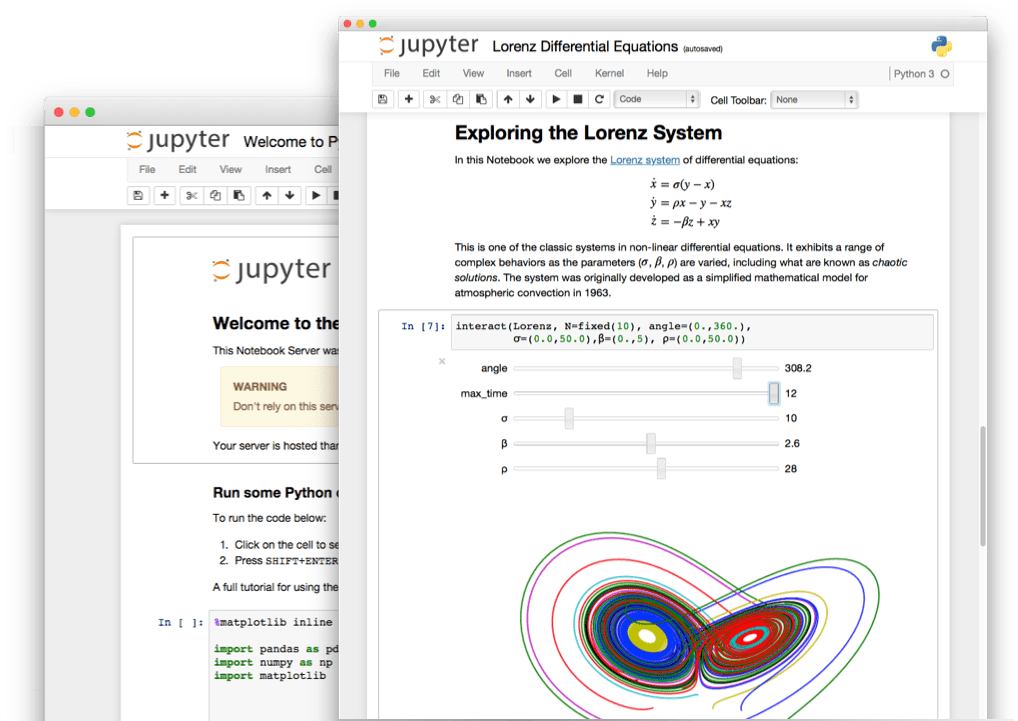
Jupyter Notebooks and Jupyter Lab
The Jupyter Notebook is an open-source web application that allows you to create and share documents that contain live code, equations, visualizations and narrative text.
JupyterLab is an interactive development environment for working with notebooks, code and data. Most importantly, JupyterLab has full support for Jupyter notebooks. Additionally, JupyterLab enables you to use text editors, terminals, data file viewers, and other custom components side by side with notebooks in a tabbed work area.
NetLogo
NetLogo is a multi-agent programmable modeling environment. It is used by tens of thousands of students, teachers and researchers worldwide.
NetLogo pageLinks of interest:
Copasi
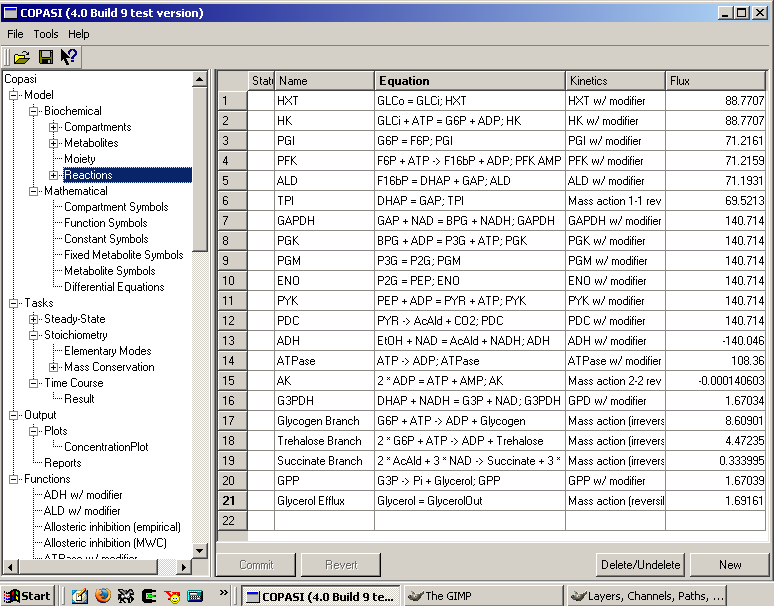
COPASI is a software application for simulation and analysis of biochemical networks and their dynamics. COPASI is a stand-alone program that supports models in the SBML standard and can simulate their behavior using ODEs or Gillespie's stochastic simulation algorithm; arbitrary discrete events can be included in such simulations.
COPASI carries out several analyses of the network and its dynamics and has extensive support for parameter estimation and optimization. COPASI provides means to visualize data in customizable plots, histograms and animations of network diagrams.
Launch Copasi Copasi pageQtOctave
From the GNU Octave Wikipedia page:
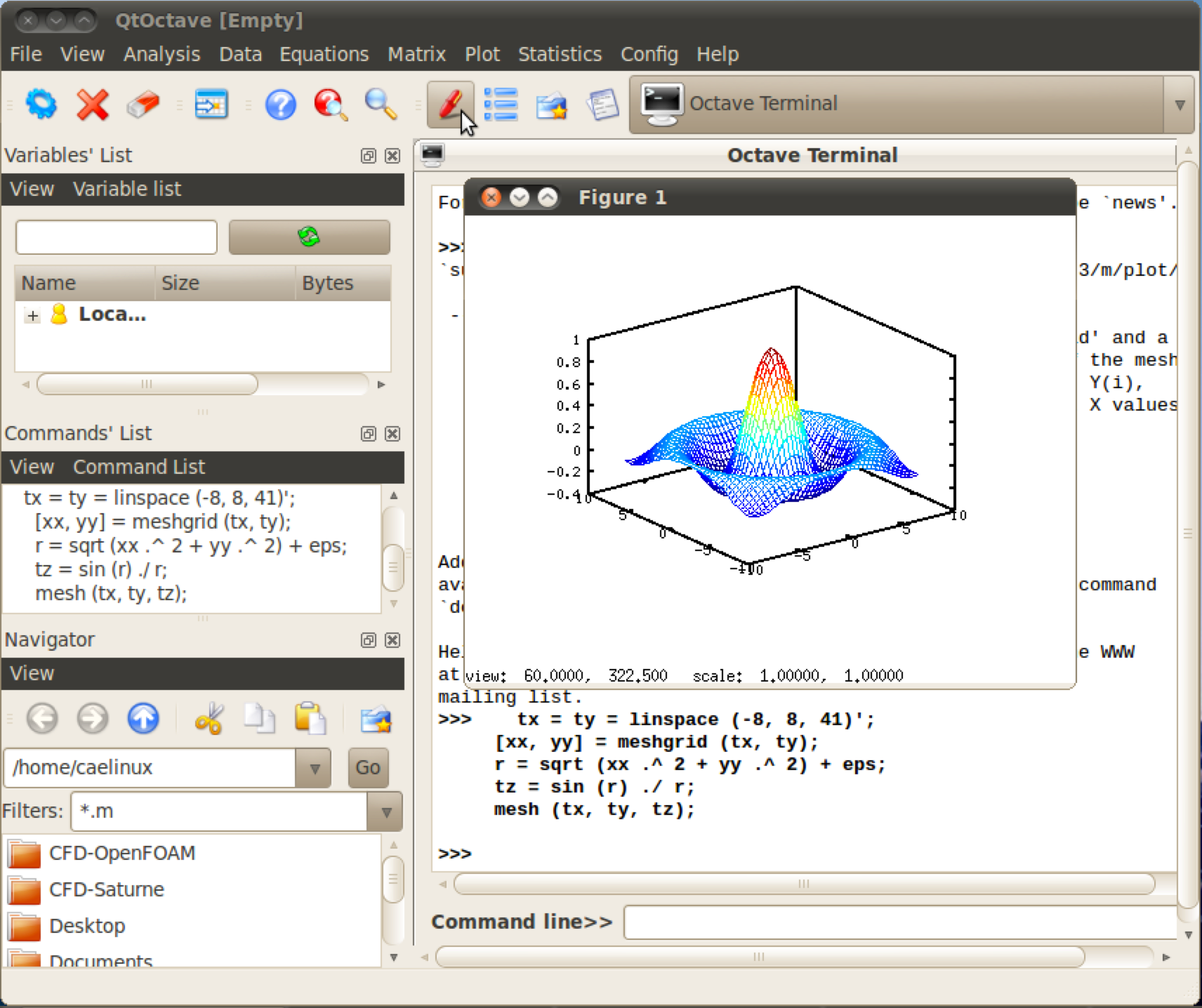
GNU Octave is a high-level programming language, primarily intended for numerical computations. It provides a command-line interface for solving linear and nonlinear problems numerically, and for performing other numerical experiments using a language that is mostly compatible with MATLAB. It may also be used as a batch-oriented language. As part of the GNU Project, it is free software under the terms of the GNU General Public License.
QtOctave is an open source GUI front-end application for GNU Octave.
ImageJ
From Introduction to ImageJ:
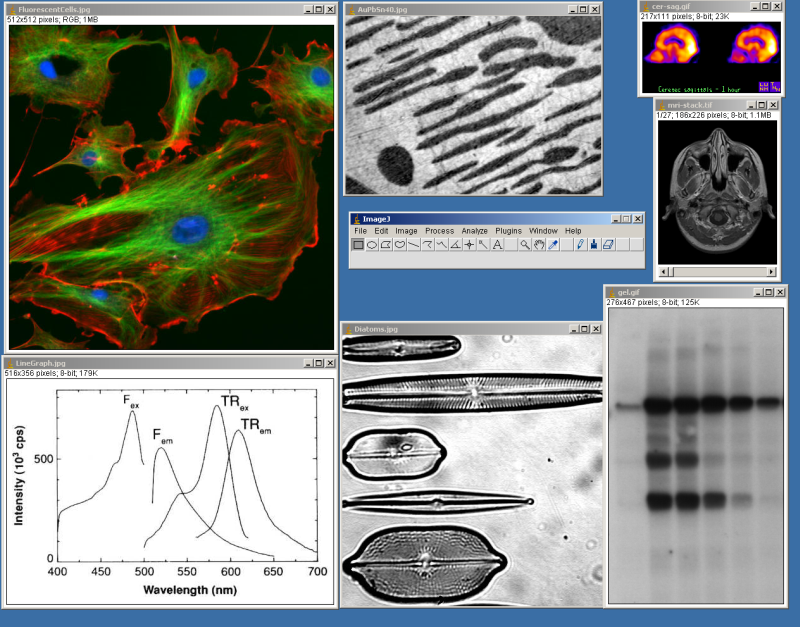
ImageJ is a public domain Java image processing program inspired by NIH Image for the Macintosh. It runs, either as an online applet or as a downloadable application, on any computer with a Java 1.4 or later virtual machine. Downloadable distributions are available for Windows, Mac OS, Mac OS X and Linux. It can display, edit, analyze, process, save and print 8-bit, 16-bit and 32-bit images. It can read many image formats including TIFF, GIF, JPEG, BMP, DICOM, FITS and "raw". It supports "stacks", a series of images that share a single window. It is multithreaded, so time-consuming operations such as image file reading can be performed in parallel with other operations.
Mesquite
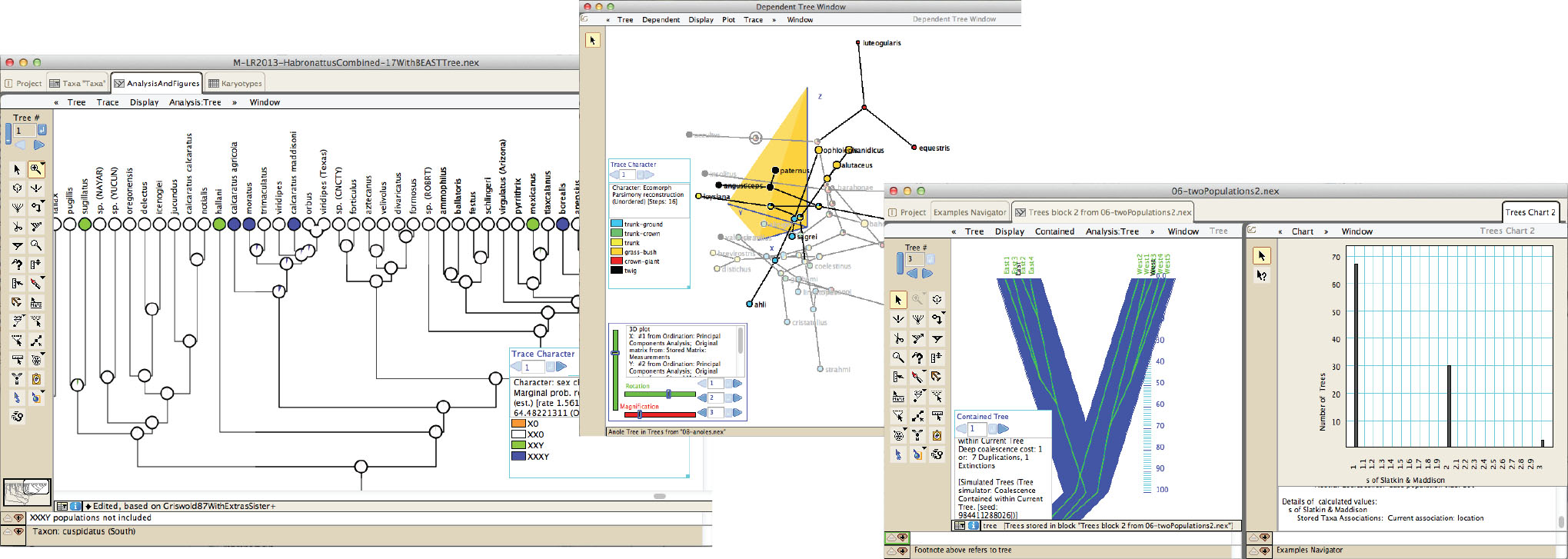
Mesquite is modular, extendible software for evolutionary biology, designed to help biologists organize and analyze comparative data about organisms. Its emphasis is on phylogenetic analysis, but some of its modules concern population genetics, while others do non-phylogenetic multivariate analysis. Because it is modular, the analyses available depend on the modules installed.
Mesquite also has many features for managing and processing data, including processing of chromatograms, sequence alignment, editing of morphometric data, and others.
PPLANE
PPLANE is a tool for phase plane analysis of a system of differential equations with 2 state variables. Users can select from a verity of numerical solvers, and control the parameters of those solvers. Solutions can be viewed in three dimensions. PPLANE also graphs the linearization about equilibrium points, displays eigenvalues, eigenvectors, nullclines, and stable & unstable orbits.
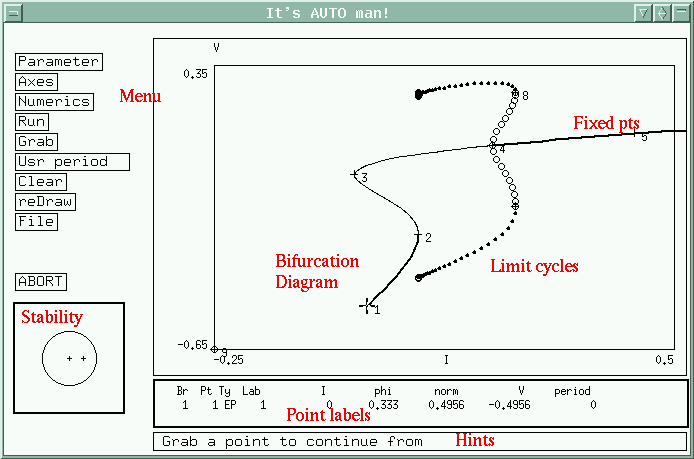
XPPAUT
XPPAUT is a general numerical tool for simulating, animating, and analyzing dynamical systems. These can range from
- discrete finite state models (e.g., McCulloch-Pitts neurons) to
- stochastic Markov models, to
- discretization of partial differential equations and integro-differential equations.
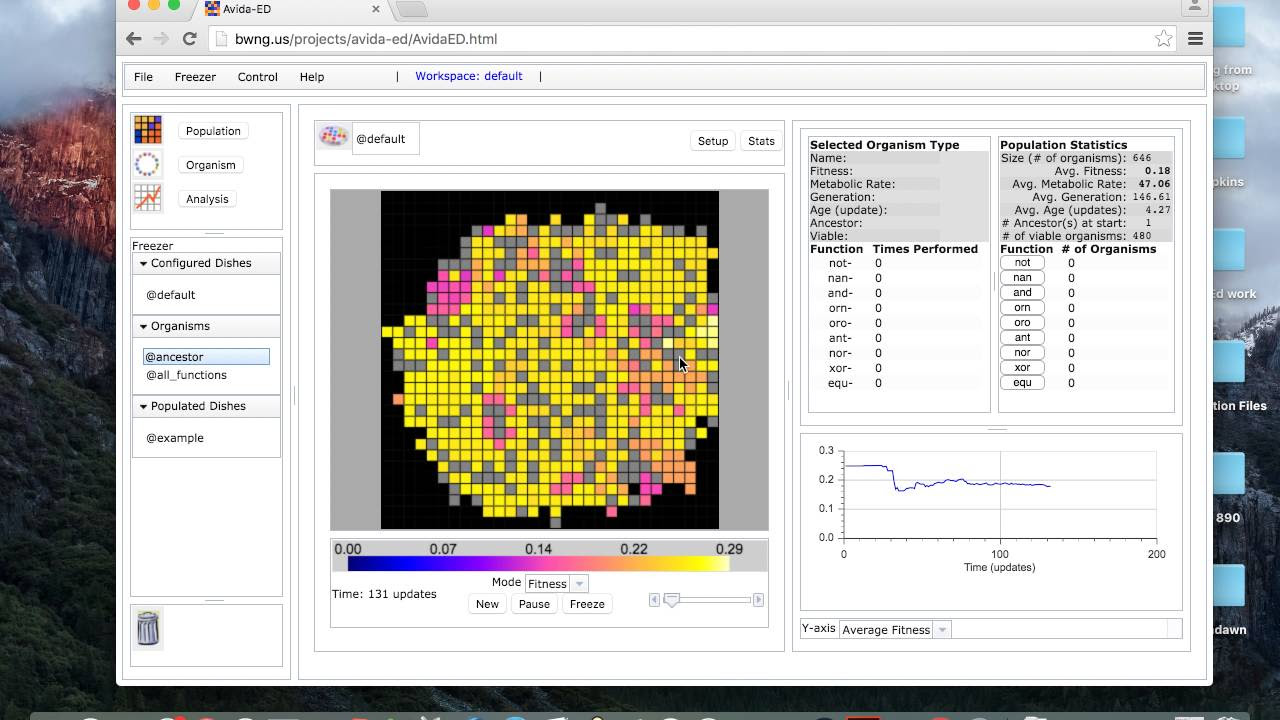
Avida-ED
Avida-ED is an award-winning educational application developed at Michigan State University for undergraduate biology courses to help students learn about evolution and scientific method by allowing them to design and perform experiments to test hypotheses about evolutionary mechanisms using evolving digital organisms.
Relevant Links
Dates: September - December, 2015
Group Description: This faculty mentoring network will provide participants with a basic introduction to teaching agent-based modeling with NetLogo. The goal is to increase faculty awareness of how NetLogo-based activities can be used to build student understanding of the modeling process and complex biology processes.
This website contains a collection of materials for teaching network-based models of the transmission of infectious diseases, including:
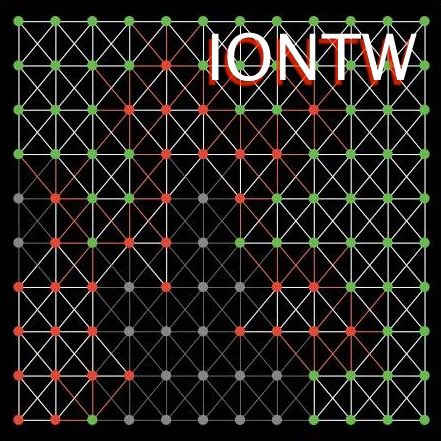
- Access to the software tool called IONTW, which stands for Infections On NeTWorks and uses the NetLogo1 programming language.
- A number of modules for active student learning at the level of advanced undergraduate and graduate students of mathematics, biology, or a related field.
- Background readings on network-based models of disease transmission.
 Using NetLogo in the Classroom
Using NetLogo in the Classroom