The purpose of this semester-long Lesson is to give students an authentic, course-based undergraduate research experience during which they learn basic and advanced microbiological and molecular biology techniques. This project begins with the isolation of a suspected Streptomyces bacterium from a soil sample and concludes with its identification. Students collect data, regarding colony and cell morphology, biochemical characteristics, the production of secondary metabolites, and employs the PCR using custom-designed primers to the Streptomyces 16s rRNA gene. The project culminates with the identification of their soil isolate using the National Center for Biotechnology Information (NCBI) web site to perform nucleotide blasts. The blastn program provides the final piece of evidence used to confirm, or not, the identification of their isolate as a Streptomyces from 16s rRNA gene sequence data, hence the title “From Dirt to Streptomyces DNA. In addition, the Lesson focuses on the Streptomyces bacteria to address several ASM aligned goals and objectives. These include prokaryotic growth phases and ways in which interactions of microorganisms among themselves and with their environment is determined by their metabolic abilities. In addition, this Lesson illustrates how microbial metabolism is important to a relevant societal issue, the need for new antibiotic discovery particularly given the rise of antibiotic resistance strains of clinically relevant bacteria. It also illustrates the microbial diversity of soil and the developmental/physiological strategies employed in such a competitive environment. This Lesson hopes to impart both the thrill and challenges associated with scientific discovery.
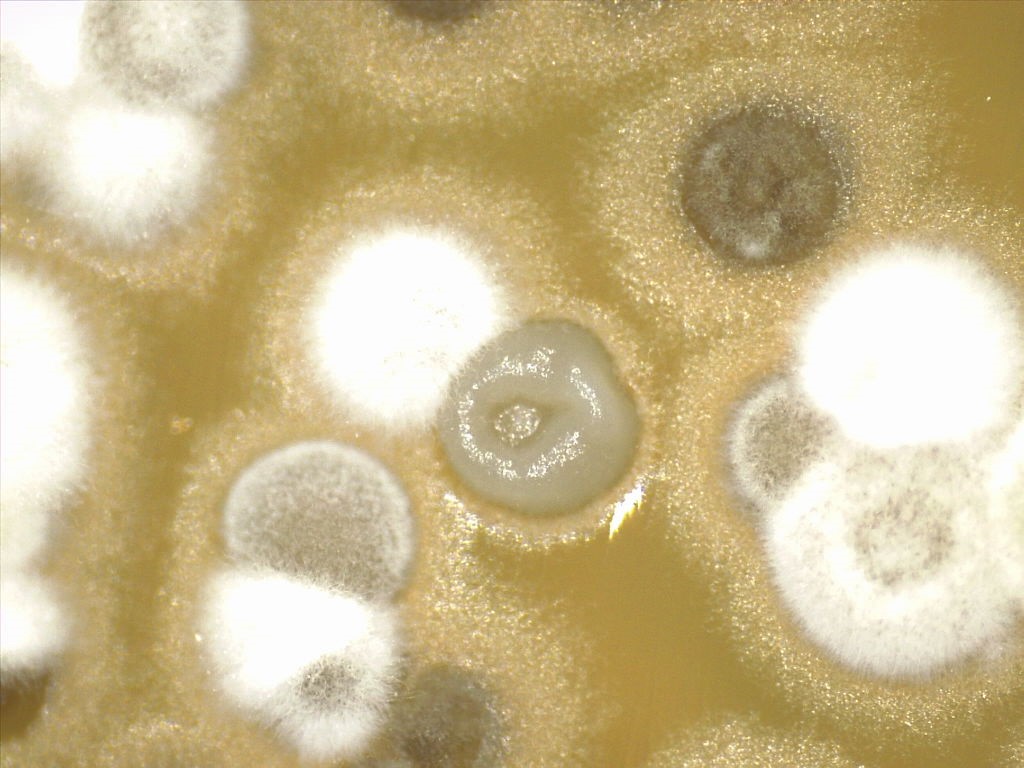
Primary image: Photomicrograph of Streptomyces colonies growing on ISP 2 agar. The Streptomyces are student isolates showing stages of morphological development. Photomicrograph by Marc A. Brodkin.

Karen Meysick onto MBIO 4873
@
on